GlycoDraw("Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc",
highlight_motif = "GlcNAc(b1-?)Man")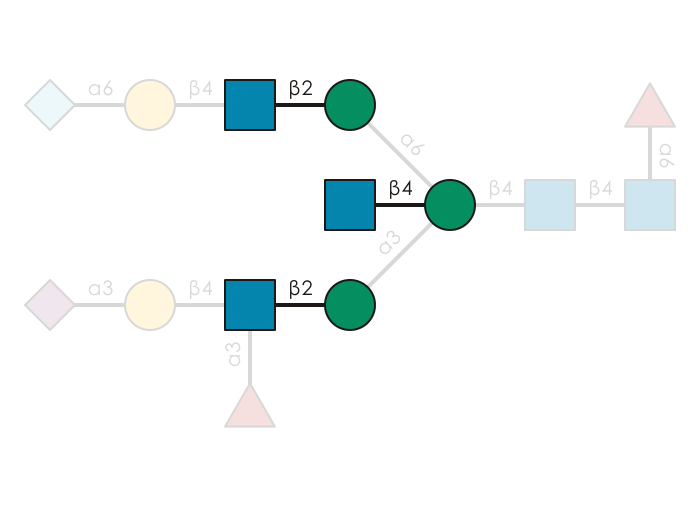
motif contains many functions to process glycans in various ways and use this processing to analyze glycans via curated motifs, graph features, and sequence features. It contains the following modules:
draw contains the GlycoDraw function to draw glycans in SNFG styleanalysis contains functions for downstream analyses of important glycan motifs etc.annotate contains functions to extract curated motifs, graph features, and sequence features from glycan sequencesgraph is used to convert glycan sequences to graphs and contains helper functions to search for motifs / check whether two sequences describe the same sequence, etc.processing contains functions to process IUPAC-condensed glycan sequences, as well as conversion functions to convert other nomenclatures into IUPAC-condensed.regex contains functionality for performing powerful regular expression-like searches on glycans; get_match is the user-facing function.query is used to interact with the databases contained in glycowork, delivering insights for sequences of interesttokenization has helper functions to map m/z–>composition, composition–>structure, structure–>motif, and moredrawing glycans in SNFG style
def GlycoDraw(
glycan:str, # IUPAC-condensed glycan sequence
vertical:bool=False, # Draw vertically
compact:bool=False, # Use compact style
show_linkage:bool=True, # Show linkage labels
dim:float=50, # Base dimension for scaling
highlight_motif:str | None=None, # Motif to highlight
highlight_termini_list:list=[], # Terminal positions (from 'terminal', 'internal', and 'flexible')
highlight_linkages:list[int] | None=None, # Which linkages to highlight in a different color; indices, starting from 0, in glycan
reverse_highlight:bool=False, # Whether to highlight everything EXCEPT highlight_motif
repeat:bool | int | str | None=None, # Repeat unit specification (True: n units, int: # of units, str: range of units)
repeat_range:list[int] | None=None, # Repeat unit range
draw_method:str | None=None, # Drawing method: None, 'chem2d', 'chem3d'
filepath:str | pathlib.Path | None=None, # Output file path
suppress:bool=False, # Suppress display
per_residue:list=[], # Per-residue intensity values (order should be the same as the monosaccharides in glycan string)
pdb_file:str | pathlib.Path | None=None, # only used when draw_method='chem3d'; already existing glycan structure
alt_text:str | None=None, # Custom ALT text for accessibility
libr:dict | None=None, # Can be modified for drawing too exotic monosaccharides
reducing_end_label:str | None=None, # Label to be drawn connected to the reducing end
restrict_vocab:bool=False, # Whether only tokens present in libr can be drawn
)->Any: # Drawing object
Renders glycan structure using SNFG symbols or chemical structure representation

def annotate_figure(
svg_input:str, # Input SVG file path
scale_range:tuple=(25, 80), # Min/max glycan dimensions
compact:bool=False, # Use compact style
glycan_size:str='medium', # Glycan size preset ('small', 'medium', 'large')
filepath:str | pathlib.Path='', # Output file path
scale_by_DE_res:pandas.DataFrame | None=None, # Differential expression results (motif_analysis.get_differential_expression)
x_thresh:float=1, # X metric threshold
y_thresh:float=0.05, # P-value threshold
x_metric:str='Log2FC', # X axis metric ('Log2FC', 'Effect size')
)->str | None: # Modified SVG code
Replaces text labels with glycan drawings in SVG figure
Creates Excel file with SNFG glycan images in a new column
downstream analyses of important glycan motifs
def get_pvals_motifs(
df:pandas.DataFrame | str, # Input dataframe or filepath (.csv/.xlsx)
glycan_col_name:str='glycan', # Column name for glycan sequences
label_col_name:str='target', # Column name for labels
zscores:bool=True, # Whether data are z-scores
thresh:float=1.645, # Threshold to separate positive/negative
sorting:bool=True, # Sort p-value dataframe
feature_set:list=['exhaustive'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
multiple_samples:bool=False, # Multiple samples with glycan columns
motifs:pandas.DataFrame | None=None, # Modified motif_list
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
)->DataFrame: # DataFrame with p-values, FDR-corrected p-values, and Cohen's d effect sizes for glycan motifs
Identifies significantly enriched glycan motifs using Welch’s t-test with FDR correction and Cohen’s d effect size calculation, comparing samples above/below threshold
glycans = ['Man(a1-3)[Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc',
'Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
'GalNAc(a1-4)GlcNAcA(a1-4)[GlcN(b1-7)]Kdo(a2-5)[Kdo(a2-4)]Kdo(a2-6)GlcOPN(b1-6)GlcOPN',
'Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
'Glc(b1-3)Glc(b1-3)Glc']
label = [3.234, 2.423, 0.733, 3.102, 0.108]
test_df = pd.DataFrame({'glycan':glycans, 'binding':label})
print("Glyco-Motif enrichment p-value test")
out = get_pvals_motifs(test_df, 'glycan', 'binding').iloc[:10,:]Glyco-Motif enrichment p-value test| motif | pval | corr_pval | effect_size | |
|---|---|---|---|---|
| 4 | GlcNAc | 0.038120 | 0.144857 | 1.530905 |
| 8 | Man | 0.054356 | 0.165364 | 1.390253 |
| 16 | Man(a1-2/3/6)Man | 0.060923 | 0.165364 | 1.308333 |
| 14 | Man(a1-3)Man | 0.034212 | 0.144857 | 1.196586 |
| 12 | Man(a1-6)Man | 0.019543 | 0.123771 | 1.168815 |
| 15 | Man(b1-4)GlcNAc | 0.019543 | 0.123771 | 1.168815 |
| 18 | GlcNAc(b1-4)GlcNAc | 0.019543 | 0.123771 | 1.168815 |
| 7 | Kdo | 0.328790 | 0.496393 | -0.811679 |
| 2 | Glc | 0.644180 | 0.679968 | -0.811679 |
| 11 | Man(a1-2)Man | 0.177461 | 0.421470 | 0.772320 |
Constructs minimal glycan structures that represent significantly enriched motifs by optimizing for motif content while minimizing structure size using subgraph isomorphism
def get_heatmap(
df:pandas.DataFrame | str | pathlib.Path, # Input dataframe or filepath (.csv/.xlsx)
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
transform:str='', # Transform data before plotting
datatype:str='response', # Data type: 'response' for quantitative values or 'presence' for presence/absence
rarity_filter:float=0.05, # Min proportion for non-zero values
filepath:str | pathlib.Path='', # Path to save plot
index_col:str='glycan', # Column to use as index
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
return_plot:bool=False, # Return plot object
show_all:bool=False, # Show all tick labels
kwargs:Any
)->tuple[typing.Any, list[str], pandas.DataFrame] | None: # None or (plot object, column names, transformed dataframe) if return_plot=True
Creates hierarchically clustered heatmap visualization of glycan/motif abundances
glycans = ['Man(a1-3)[Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc',
'Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
'GalNAc(a1-4)GlcNAcA(a1-4)[GlcN(b1-7)]Kdo(a2-5)[Kdo(a2-4)]Kdo(a2-6)GlcN4P(b1-6)GlcN4P',
'Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
'Glc(b1-3)Glc(b1-3)Glc']
label = [3.234, 2.423, 0.733, 3.102, 0.108]
label2 = [0.134, 0.345, 1.15, 0.233, 2.981]
label3 = [0.334, 0.245, 1.55, 0.133, 2.581]
test_df = pd.DataFrame([label, label2, label3], columns = glycans)
get_heatmap(test_df, motifs = True, feature_set = ['known', 'exhaustive'])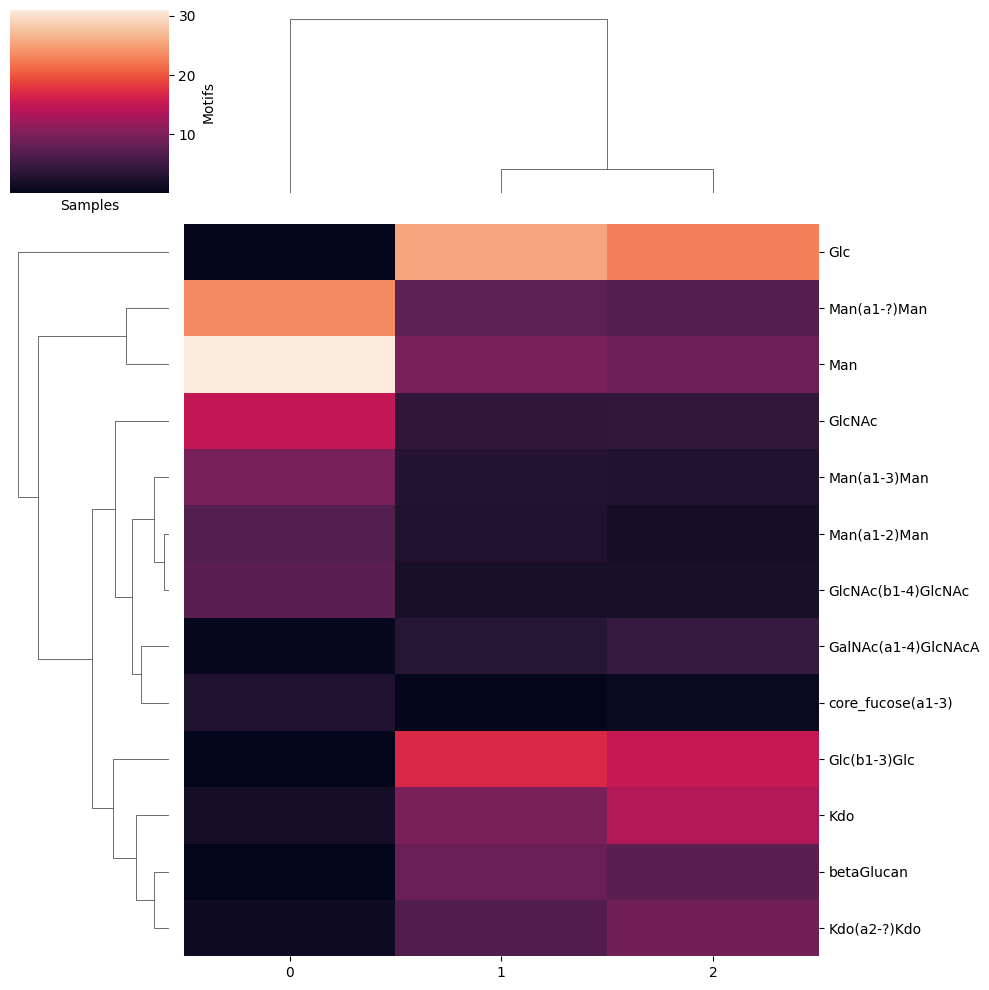
def plot_embeddings(
glycans:list, # List of IUPAC-condensed glycan sequences
emb:dict[str, numpy.ndarray] | pandas.DataFrame | None=None, # Glycan embeddings dict/DataFrame; defaults to SweetNet embeddings
label_list:list[typing.Any] | None=None, # Labels for coloring points
shape_feature:str | None=None, # Monosaccharide/bond for point shapes
filepath:str | pathlib.Path='', # Path to save plot
alpha:float=0.8, # Point transparency
palette:str='colorblind', # Color palette for groups
kwargs:Any
)->None: # Keyword args passed to seaborn scatterplot
Visualizes learned glycan embeddings using t-SNE dimensionality reduction with optional group coloring
Download completed.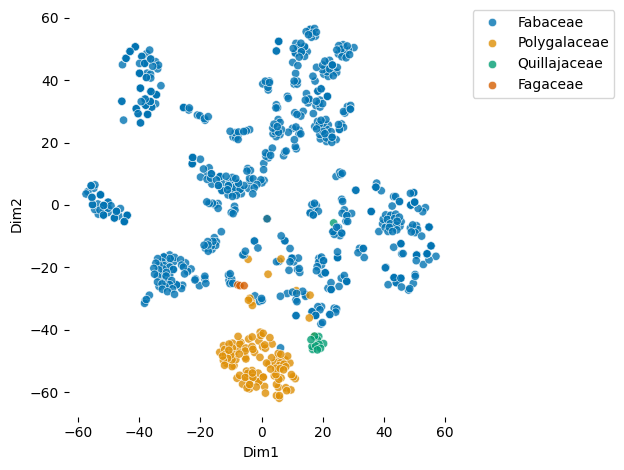
def characterize_monosaccharide(
sugar:str, # Monosaccharide or linkage to analyze
df:pandas.DataFrame | None=None, # DataFrame with glycan column 'glycan'; defaults to df_species
mode:str='sugar', # Analysis mode: 'sugar', 'bond', 'sugarbond'
glycan_col_name:str='glycan', # Column name for glycan sequences
rank:str | None=None, # Column name for group filtering
focus:str | None=None, # Row value for group filtering
modifications:bool=False, # Consider modified monosaccharides
filepath:str | pathlib.Path='', # Path to save plot
thresh:int=10, # Minimum count threshold for inclusion
)->None:
Analyzes connectivity and modification patterns of specified monosaccharides/linkages in glycan sequences
def get_differential_expression(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (col 1) and abundance values in subsequent columns
group1:list, # Column indices/names for first group
group2:list, # Column indices/names for second group
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['exhaustive', 'known'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
paired:bool=False, # Whether samples are paired
impute:bool=True, # Replace zeros with Random Forest model
sets:bool=False, # Identify clusters of correlated glycans
set_thresh:float=0.9, # Correlation threshold for clusters
effect_size_variance:bool=False, # Calculate effect size variance
min_samples:float=0.1, # Min percent of non-zero samples required
grouped_BH:bool=False, # Use two-stage adaptive Benjamini-Hochberg
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float | dict=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
glycoproteomics:bool=False, # Whether data is from glycoproteomics
level:str='peptide', # Analysis level for glycoproteomics
monte_carlo:bool=False, # Use Monte Carlo for technical variation
random_state:int | numpy.random._generator.Generator | None=None, # optional random state for reproducibility
)->DataFrame: # DataFrame with log2FC, p-values, FDR-corrected p-values, and Cohen's d/Mahalanobis distance effect sizes
Performs differential expression analysis using Welch’s t-test (or Hotelling’s T2 for sets) with multiple testing correction on glycomics abundance data
You're working with an alpha of 0.044390023979542614 that has been adjusted for your sample size of 40.
Significance inflation detected. The CLR/ALR transformation possibly cannot handle this dataset. Consider running again with a higher gamma value. Proceed with caution; for now switching to Bonferroni correction to be conservative about this.| Glycan | Mean abundance | Log2FC | p-val | corr p-val | significant | corr Levene p-val | Effect size | Equivalence p-val | |
|---|---|---|---|---|---|---|---|---|---|
| 6 | Gal | 18.570780 | -2.810982 | 3.038799e-22 | 4.558198e-21 | True | 3.684477e-05 | -12.047275 | 1.000000 |
| 11 | Neu5Ac(a2-3)Gal | 12.364384 | -2.168027 | 3.371284e-22 | 5.056926e-21 | True | 6.854560e-08 | -11.981208 | 1.000000 |
| 14 | Gal(b1-3)GalNAc | 12.746417 | -2.286370 | 9.062568e-22 | 1.359385e-20 | True | 3.088705e-05 | -11.369661 | 1.000000 |
| 10 | Neu5Ac | 16.580453 | -2.632591 | 3.241244e-21 | 4.861866e-20 | True | 1.631811e-05 | -10.626734 | 1.000000 |
| 7 | GalNAc | 12.906669 | -2.316123 | 8.372261e-20 | 1.255839e-18 | True | 1.783496e-05 | -8.940591 | 1.000000 |
| 13 | Neu5Ac(a2-8)Neu5Ac | 0.038743 | 6.249081 | 1.051533e-16 | 1.577300e-15 | True | 9.004822e-06 | 6.102326 | 1.000000 |
| 4 | Oglycan_core1 | 7.969780 | -1.578350 | 1.959493e-14 | 2.939239e-13 | True | 7.682079e-06 | -4.593089 | 1.000000 |
| 8 | GalOS | 0.160252 | 3.935439 | 9.315000e-14 | 1.397250e-12 | True | 3.550473e-04 | 4.215097 | 1.000000 |
| 0 | H_antigen_type2 | 0.247550 | 3.251957 | 8.838338e-13 | 1.325751e-11 | True | 9.729411e-06 | 3.718853 | 1.000000 |
| 5 | Mucin_elongated_core2 | 4.776637 | -0.916792 | 1.468039e-08 | 2.202059e-07 | True | 3.997627e-06 | -2.096764 | 1.000000 |
| 9 | GlcNAc6S(b1-6)GalNAc | 1.047725 | 2.101894 | 6.500297e-08 | 9.750445e-07 | True | 7.342527e-05 | 1.905049 | 1.000000 |
| 12 | Neu5Ac(a2-6)GalNAc | 4.017075 | -0.636913 | 1.990078e-05 | 2.985117e-04 | True | 9.729411e-06 | -1.258696 | 1.000000 |
| 1 | Internal_LacNAc_type2 | 2.332012 | 0.446722 | 1.687358e-04 | 2.531038e-03 | True | 3.997627e-06 | 1.043131 | 1.000000 |
| 3 | Disialyl_T_antigen | 3.796897 | -0.550242 | 2.034837e-04 | 3.052256e-03 | True | 2.044072e-06 | -1.024620 | 1.000000 |
| 2 | Terminal_LacNAc_type2 | 2.444625 | -0.100778 | 5.115432e-01 | 1.000000e+00 | False | 1.828193e-06 | -0.149593 | 0.514385 |
def get_volcano(
df_res:pandas.DataFrame | str | pathlib.Path, # DataFrame from get_differential_expression with columns [Glycan, Log2FC, p-val, corr p-val]
y_thresh:float=0.05, # Corrected p threshold for labeling
x_thresh:float=0, # Absolute x metric threshold for labeling
n:int | None=None, # Sample size for Bayesian-Adaptive Alpha
label_changed:bool=True, # Add text labels to significant points
x_metric:str='Log2FC', # x-axis metric: 'Log2FC' or 'Effect size'
annotate_volcano:bool=False, # Annotate dots with SNFG images
filepath:str='', # Path to save plot
kwargs:Any
)->None: # Displays volcano plot
Creates volcano plot showing -log10(FDR-corrected p-values) vs Log2FC or effect size
You're working with a default alpha of 0.05. Set sample size (n = ...) for Bayesian-Adaptive Alpha Adjustment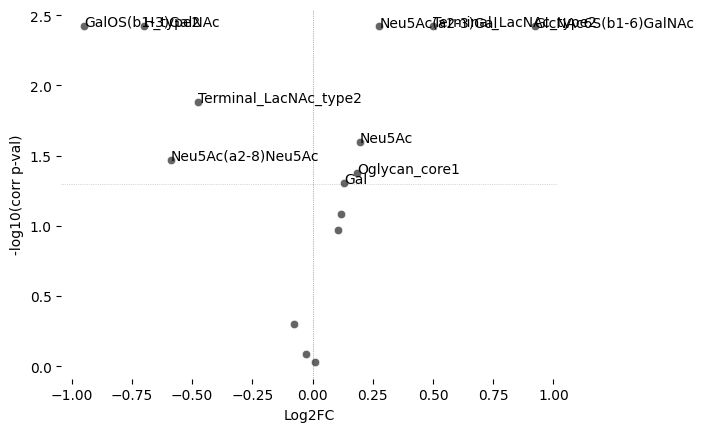
Visualizes glycan detection frequency across samples with intensity-based ordering
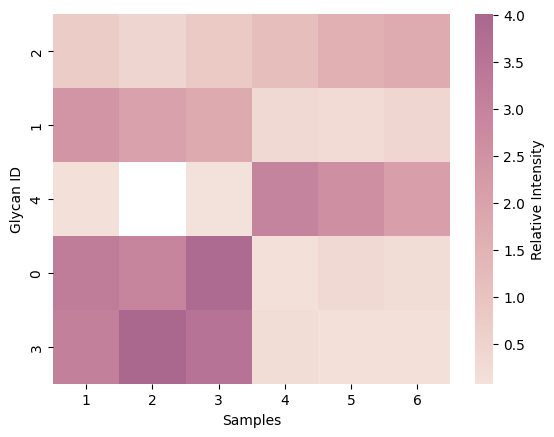
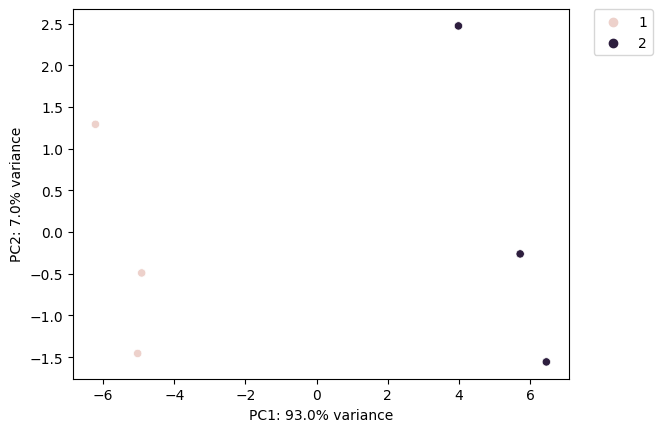
def get_pca(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (col 1), abundances in columns
groups:list[int] | pandas.DataFrame | None=None, # Group labels (e.g., [1,1,1,2,2,2,3,3,3]) or metadata DataFrame with 'id' column
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known', 'exhaustive'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
pc_x:int=1, # Principal component for x-axis
pc_y:int=2, # Principal component for y-axis
color:str | None=None, # Column in metadata for color grouping; recommended to be categorical
shape:str | None=None, # Column in metadata for shape grouping; recommended to be categorical
size:str | None=None, # Column in metadata for point size control; recommended to be scalar
filepath:str | pathlib.Path='', # Path to save plot
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
rarity_filter:float=0.05, # Min proportion for non-zero values
)->None:
Performs PCA on glycan/motif abundance data with group-based visualization
Creates histogram of p-values from differential expression analysis
Generates MA plot (mean abundance vs log2 fold change) from differential expression results
def get_glycanova(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (col 1) and abundance values in columns
groups:list, # Group labels for samples (e.g., [1,1,1,2,2,2,3,3,3])
impute:bool=True, # Replace zeros with Random Forest model
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['exhaustive', 'known'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
min_samples:float=0.1, # Min percent of non-zero samples required
posthoc:bool=True, # Perform Tukey's HSD test post-hoc
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
random_state:int | numpy.random._generator.Generator | None=None, # optional random state for reproducibility
)->tuple: # (ANOVA results with F-stats and omega-squared effect sizes, post-hoc results)
Performs one-way ANOVA with omega-squared effect size calculation and optional Tukey’s HSD post-hoc testing on glycomics data across multiple groups
You're working with an alpha of 0.06364810000741428 that has been adjusted for your sample size of 12.| Glycan | F statistic | p-val | corr p-val | significant | Effect size | |
|---|---|---|---|---|---|---|
| 4 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 9.538792 | 0.005977 | 0.041838 | True | 0.448494 |
| 7 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Neu5Ac(a2-... | 6.706481 | 0.016476 | 0.057665 | True | -0.052876 |
| 0 | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 2.725019 | 0.118763 | 0.169985 | False | 0.141106 |
| 2 | Neu5Ac(a2-3)Gal(b1-3)GalNAc | 2.324103 | 0.153551 | 0.169985 | False | 0.111983 |
| 6 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]Ga... | 2.676719 | 0.122402 | 0.169985 | False | 0.137699 |
| 5 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3/6)[GlcNAc(b1-... | 2.159181 | 0.171420 | 0.171420 | False | 0.099422 |
| 3 | Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Ga... | 2.086710 | 0.180073 | 0.180073 | False | 0.093789 |
| 1 | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 1.976577 | 0.194268 | 0.194268 | False | 0.085093 |
| 8 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)... | 0.000000 | 1.000000 | 1.000000 | False | 0.352111 |
def get_meta_analysis(
effect_sizes:numpy.ndarray | list[float], # List of Cohen's d/other effect sizes
variances:numpy.ndarray | list[float], # Associated variance estimates
model:str='fixed', # 'fixed' or 'random' effects model
filepath:str='', # Path to save Forest plot
study_names:list=[], # Names corresponding to each effect size
)->tuple: # (combined effect size, two-tailed p-value)
Performs fixed/random effects meta-analysis using DerSimonian-Laird method for between-study variance estimation, with optional Forest plot visualization
(-5.326913553837341, 3.005077298112724e-09)
def get_time_series(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with sample IDs as 'sampleID_timepoint_replicate' in col 1 (e.g., T1_h5_r1)
impute:bool=True, # Replace zeros with Random Forest model
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known', 'exhaustive'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
degree:int=1, # Polynomial degree for regression
min_samples:float=0.1, # Min percent of non-zero samples required
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float | dict=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
)->DataFrame: # DataFrame with regression coefficients and FDR-corrected p-values
Analyzes time series glycomics data using polynomial regression
t_dic = {}
t_dic["ID"] = ["D1_h5_r1", "D1_h5_r2", "D1_h5_r3", "D1_h10_r1", "D1_h10_r2", "D1_h10_r3", "D1_h15_r1", "D1_h15_r2", "D1_h15_r3"]
t_dic["Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc"] = [0.33, 0.31, 0.35, 1.51, 1.57, 1.66, 2.11, 2.04, 2.09]
t_dic["Fuc(a1-2)Gal(b1-3)GalNAc"] = [0.78, 1.01, 0.98, 0.88, 1.11, 0.72, 1.22, 1.00, 0.54]
t_dic["Neu5Ac(a2-6)GalNAc"] = [0.11, 0.09, 0.14, 0.02, 0.07, 0.10, 0.11, 0.09, 0.08]
get_time_series(pd.DataFrame(t_dic).set_index("ID").T)You're working with an alpha of 0.0694557066556809 that has been adjusted for your sample size of 9.| Glycan | Change | p-val | corr p-val | significant | |
|---|---|---|---|---|---|
| 0 | Fuc(a1-2)Gal(b1-3)GalNAc | -0.006923 | 0.202954 | 0.202954 | False |
| 1 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]Ga... | 0.019568 | 0.091398 | 0.202954 | False |
| 2 | Neu5Ac(a2-6)GalNAc | -0.013189 | 0.160749 | 0.202954 | False |
def get_jtk(
df_in:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (first column), then groups arranged by ascending timepoints
timepoints:int, # Number of timepoints (each must have same number of replicates)
interval:int, # Time units between experimental timepoints
periods:list=[12, 24], # Timepoints per cycle to test
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known', 'exhaustive', 'terminal'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
correction_method:str='two-stage', # Multiple testing correction method
)->DataFrame: # DataFrame with JTK results: adjusted p-values, period length, lag phase, amplitude
Identifies rhythmically expressed glycans using Jonckheere-Terpstra-Kendall algorithm for time series analysis
t_dic = {}
t_dic["Neu5Ac(a2-3)Gal(b1-3)GalNAc"] = [0.433138901, 0.149729209, 0.358018822, 0.537641256, 1.526963756, 1.349986672, 0.75156406, 0.736710183]
t_dic["Gal(b1-3)GalNAc"] = [0.919762334, 0.760237184, 0.725566662, 0.459945797, 0.523801515, 0.695106926, 0.627632047, 1.183511209]
t_dic["Gal(b1-3)[Neu5Ac(a2-6)]GalNAc"] = [0.533138901, 0.119729209, 0.458018822, 0.637641256, 1.726963756, 1.249986672, 0.55156406, 0.436710183]
t_dic["Fuc(a1-2)Gal(b1-3)GalNAc"] = [3.862169504, 5.455032837, 3.858163289, 5.614650335, 3.124254095, 4.189550337, 4.641831312, 4.19538484]
tps = 8 # number of timepoints in experiment
periods = [8] # potential cycles to test
interval = 3 # units of time between experimental timepoints
t_df = pd.DataFrame(t_dic).T
t_df.columns = ["T3", "T6", "T9", "T12", "T15", "T18", "T21", "T24"]
get_jtk(t_df.reset_index(), tps, interval, periods = periods)You're working with an alpha of 0.22004505213567527 that has been adjusted for your sample size of 1.| Molecule_Name | Adjusted_P_value | Period_Length | Lag_Phase | Amplitude | significant | |
|---|---|---|---|---|---|---|
| 0 | Gal(b1-3)GalNAc | 0.005945 | 8 | 12 | 0.928571 | True |
| 1 | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 0.053172 | 8 | 12 | 0.642857 | True |
| 2 | Neu5Ac(a2-3)Gal(b1-3)GalNAc | 0.063487 | 8 | 12 | 0.571429 | True |
| 3 | Fuc(a1-2)Gal(b1-3)GalNAc | 0.265510 | 8 | 3 | 0.357143 | False |
You're working with an alpha of 0.22004505213567527 that has been adjusted for your sample size of 1.| Molecule_Name | Adjusted_P_value | Period_Length | Lag_Phase | Amplitude | significant | |
|---|---|---|---|---|---|---|
| 0 | Terminal_Neu5Ac(a2-3/6) | 0.014062 | 8 | 12 | 0.785714 | True |
| 1 | Terminal_Neu5Ac(a2-6) | 0.014062 | 8 | 12 | 0.785714 | True |
| 2 | Terminal_Neu5Ac(a2-3) | 0.107762 | 8 | 9 | 0.500000 | True |
| 3 | Terminal_Fuc(a1-2) | 0.265510 | 8 | 3 | 0.357143 | False |
| 4 | Terminal_Gal(b1-3) | 0.265510 | 8 | 3 | 0.357143 | False |
def get_biodiversity(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (col 1), abundances in columns
group1:list, # First group column indices or group labels
group2:list, # Second group indices or additional group labels
metrics:list=['alpha', 'beta'], # Diversity metrics to calculate
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['exhaustive', 'known'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
paired:bool=False, # Whether samples are paired
permutations:int=999, # Number of permutations for ANOSIM/PERMANOVA
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float | dict=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
random_state:int | numpy.random._generator.Generator | None=None, # optional random state for reproducibility
)->tuple: # First DataFrame with diversity indices and test statistics, second with beta-diversity distance matrix
Calculates alpha (Shannon/Simpson) and beta (ANOSIM/PERMANOVA) diversity measures from glycomics data
You're working with an alpha of 0.044390023979542614 that has been adjusted for your sample size of 40.( Metric Group1 mean Group2 mean p-val \
0 Beta diversity (ANOSIM) NaN NaN 0.000000
1 Beta diversity (PERMANOVA) NaN NaN 0.000000
2 simpson_diversity 0.876445 0.874168 0.000520
3 shannon_diversity 2.242778 2.224732 0.001402
4 species_richness 15.000000 15.000000 1.000000
Effect size corr p-val significant
0 1.000000 0.000000 True
1 263.321829 0.000000 True
2 -0.932303 0.000520 True
3 -0.835249 0.001402 True
4 0.000000 1.000000 False ,
array([[ 0. , 2.21397052, 3.06333292, ..., 11.24499348,
11.24499348, 11.24499348],
[ 2.21397052, 0. , 2.31028564, ..., 9.50451026,
9.50451026, 9.50451026],
[ 3.06333292, 2.31028564, 0. , ..., 10.63189474,
10.63189474, 10.63189474],
...,
[11.24499348, 9.50451026, 10.63189474, ..., 0. ,
0. , 0. ],
[11.24499348, 9.50451026, 10.63189474, ..., 0. ,
0. , 0. ],
[11.24499348, 9.50451026, 10.63189474, ..., 0. ,
0. , 0. ]]))
def get_SparCC(
df1:pandas.DataFrame | str | pathlib.Path, # First DataFrame with glycans in rows (col 1) and abundances in columns
df2:pandas.DataFrame | str | pathlib.Path, # Second DataFrame with same format as df1
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known', 'exhaustive'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
partial_correlations:bool=False, # Use regularized partial correlations
)->tuple: # (Spearman correlation matrix, FDR-corrected p-value matrix)
Calculates SparCC (Sparse Correlations for Compositional Data) between two matching datasets (e.g., glycomics)
df1 = glycomics_data_loader.time_series_N_PMID32149347
df2 = glycomics_data_loader.time_series_O_PMID32149347
df1 = pd.merge(df1, df2[['ID']], on = 'ID', how = 'inner')
df2 = pd.merge(df2, df1[['ID']], on = 'ID', how = 'inner')
df1 = df1.set_index(df1.columns.tolist()[0]).T.reset_index()
df2 = df2.set_index(df2.columns.tolist()[0]).T.reset_index()
corr, pval = get_SparCC(df1, df2, motifs = True, transform = "CLR")
sns.clustermap(corr)You're working with an alpha of 0.04787928055709467 that has been adjusted for your sample size of 31.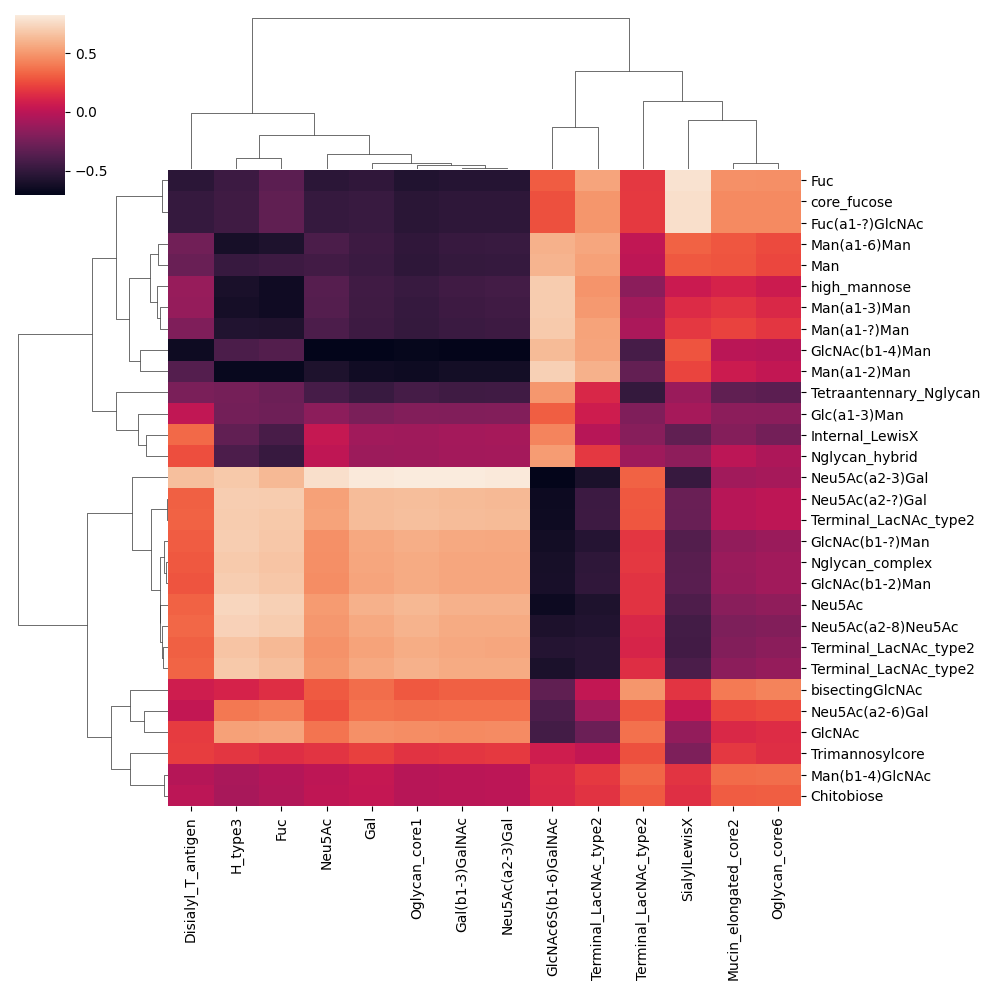
def get_roc(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with glycans in rows (col 1), abundances in columns
group1:list, # First group indices/names
group2:list, # Second group indices/names
motifs:bool=False, # Analyze motifs instead of sequences
feature_set:list=['known', 'exhaustive'], # Feature sets to use; exhaustive, known, terminal1, terminal2, terminal3, chemical, graph, custom, size_branch
paired:bool=False, # Whether samples are paired
impute:bool=True, # Replace zeros with Random Forest model
min_samples:float=0.1, # Min percent of non-zero samples required
custom_motifs:list=[], # Custom motifs if using 'custom' feature set
transform:str | None=None, # Transformation type: "CLR" or "ALR"
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float | dict=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
filepath:str | pathlib.Path='', # Path to save ROC plot
multi_score:bool=False, # Find best multi-glycan score
random_state:int | numpy.random._generator.Generator | None=None, # optional random state for reproducibility
)->list[tuple[str, float]] | dict[typing.Any, tuple[str, float]] | tuple[sklearn.linear_model._logistic.LogisticRegression, float]: # (Feature scores with ROC AUC values)
Calculates ROC curves and AUC scores for glycans/motifs or multi-glycan classifiers
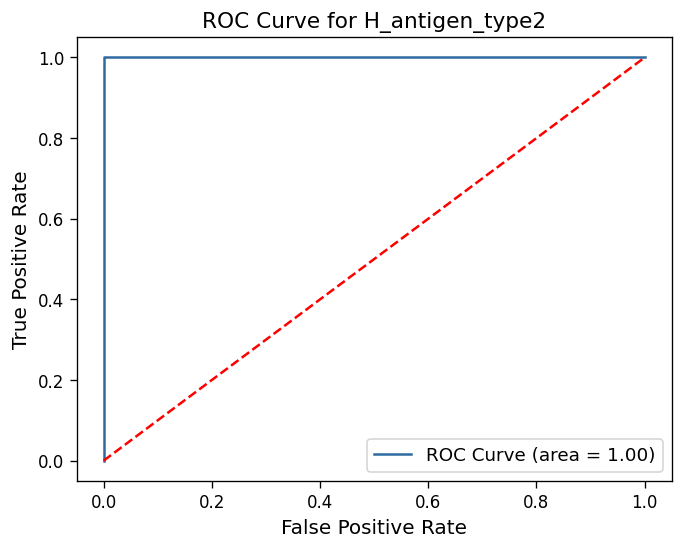
[('H_antigen_type2', 1.0),
('GalOS', 1.0),
('GlcNAc6S(b1-6)GalNAc', 1.0),
('Neu5Ac(a2-8)Neu5Ac', 1.0),
('Internal_LacNAc_type2', 0.85),
('Terminal_LacNAc_type2', 0.5),
('Neu5Ac(a2-6)GalNAc', 0.15000000000000002),
('Disialyl_T_antigen', 0.09999999999999998),
('Oglycan_core1', 0.0),
('Mucin_elongated_core2', 0.0),
('Gal', 0.0),
('GalNAc', 0.0),
('Neu5Ac', 0.0),
('Neu5Ac(a2-3)Gal', 0.0),
('Gal(b1-3)GalNAc', 0.0)]
def get_lectin_array(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with samples as rows and lectins as columns, first column containing sample IDs
group1:list, # First group indices/names
group2:list, # Second group indices/names
paired:bool=False, # Whether samples are paired
transform:str='', # Optional log2 transformation
)->DataFrame: # DataFrame with altered glycan motifs, supporting lectins, and effect sizes
Analyzes lectin microarray data by mapping lectin binding patterns to glycan motifs, calculating Cohen’s d effect sizes between groups and clustering results by significance
Lectin "Ab-LeB-1" is not found in our annotated lectin library and is excluded from analysis.
Lectin "APA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "APP" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Blood Group B [CLCP-19B]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Blood Group H2" is not found in our annotated lectin library and is excluded from analysis.
Lectin "CA19-9 [121SLE]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "CCA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "CD15 [ICRF29-2]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "CD15 [MY-1]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "CD15 [SP-159]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Forssman" is not found in our annotated lectin library and is excluded from analysis.
Lectin "IAA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "IRA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Le X [P12]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Lewis A [7LE]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Lewis B [218]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "Lewis Y [F3]" is not found in our annotated lectin library and is excluded from analysis.
Lectin "LFA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "LPA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "MNA-M " is not found in our annotated lectin library and is excluded from analysis.
Lectin "MUC5Ac Ab" is not found in our annotated lectin library and is excluded from analysis.
Lectin "PMA" is not found in our annotated lectin library and is excluded from analysis.
Lectin "PTA_1" is not found in our annotated lectin library and is excluded from analysis.
Lectin "PTA_2" is not found in our annotated lectin library and is excluded from analysis.
Lectin "SNA-S" is not found in our annotated lectin library and is excluded from analysis.
Lectin "SNA-V" is not found in our annotated lectin library and is excluded from analysis.
Lectin "VFA" is not found in our annotated lectin library and is excluded from analysis.| motif | named_motifs | lectin(s) | change | score | significance | |
|---|---|---|---|---|---|---|
| 39 | Neu5Ac(a2-6)Gal(b1-3)GlcNAc | [Internal_LacNAc_type1] | PSL, SNA, TJA-I, BDA, BPA, WGA_1, WGA_2 | down | 11.32 | highly significant |
| 38 | Neu5Ac(a2-6)Gal(b1-4)GlcNAc | [Internal_LacNAc_type2] | PSL, SNA, TJA-I, BDA, BPA, ECA, RCA120, Ricin ... | down | 10.81 | highly significant |
| 7 | Man(a1-2) | [] | ASA, Con A, CVN, HHL, SVN_1, GRFT, SVN_2, SNA-... | up | 4.83 | moderately significant |
| 14 | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc... | [Chitobiose, Trimannosylcore, Terminal_LacNAc_... | CA, CAA, DSA_1, DSA_2, DSA_3, AMA, BDA, BPA, C... | up | 3.51 | moderately significant |
| 4 | Gal(b1-3)GalNAc | [Oglycan_core1] | ACA, AIA, MPA, PNA_1, PNA_2, BDA, BPA | up | 3.48 | moderately significant |
| 43 | Neu5Ac(a2-6)GalNAc(b1-4)GlcNAc | [Internal_LacdiNAc_type2] | SNA, CSA, SBA, VVA_1, VVA_2, WFA, BPA, ECA, ST... | down | 2.86 | moderately significant |
| 10 | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][G... | [Chitobiose, Trimannosylcore, Terminal_LacNAc_... | Blackbean, Calsepa, PHA-E_1, PHA-E_2, AMA, BDA... | up | 2.70 | moderately significant |
| 16 | Fuc(a1-2)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Ga... | [Internal_LacNAc_type2, H_type3] | Cholera Toxin, AAA, AAL, ACA, AIA, AOL, BDA, B... | up | 2.51 | moderately significant |
| 15 | Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Gl... | [Internal_LacNAc_type2] | Cholera Toxin, ACA, AIA, BDA, BPA, CSA, ECA, L... | up | 2.46 | moderately significant |
| 47 | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Ma... | [Chitobiose, Trimannosylcore, core_fucose] | TL, AAL, AMA, AOL, Con A, GNA, GNL, HHL, LcH, ... | up | 2.36 | moderately significant |
| 18 | Man(a1-6) | [] | Con A, GNA, GNL, HHL, NPA, SNA-II, UDA | up | 2.30 | moderately significant |
| 17 | Man(a1-3) | [] | Con A, GNA, GNL, HHL, NPA, SNA-II, UDA | up | 2.30 | moderately significant |
| 22 | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Ma... | [Chitobiose, Trimannosylcore, Terminal_LacNAc_... | DSA_1, DSA_2, DSA_3, AMA, BDA, Blackbean, BPA,... | up | 2.05 | moderately significant |
| 46 | Fuc(a1-2)Gal(b1-3)GalNAc | [H_type3, Oglycan_core1] | TJA-II, AAA, AAL, ACA, AIA, AOL, BDA, BPA, MPA... | up | 1.96 | moderately significant |
| 3 | Fuc(a1-6) | [] | AAL, AOL, LcH, PSA | up | 1.70 | moderately significant |
| 34 | Neu5Ac(a2-3)Gal(b1-3)GalNAc | [Oglycan_core1] | MAL-II, ACA, AIA, BDA, BPA, MPA, PNA_1, PNA_2,... | up | 1.59 | moderately significant |
| 6 | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | [Chitobiose, Trimannosylcore] | AMA, Con A, GNA, GNL, HHL, NPA, SNA-II, UDA, W... | up | 1.58 | moderately significant |
| 11 | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6... | [Chitobiose, Trimannosylcore] | Blackbean, PHA-L, AMA, Con A, GNA, GNL, HHL, N... | up | 1.44 | moderately significant |
| 42 | GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-... | [Chitobiose, Trimannosylcore, bisectingGlcNAc] | RPA, AMA, Blackbean, Con A, GNA, GNL, HHL, NPA... | up | 1.40 | moderately significant |
| 41 | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-... | [Chitobiose, Trimannosylcore, bisectingGlcNAc] | RPA, AMA, Con A, GNA, GNL, HHL, NPA, SNA-II, U... | up | 1.36 | moderately significant |
| 23 | Gal(b1-4)GlcNAc | [Terminal_LacNAc_type2] | ECA, RCA120, Ricin B Chain, SJA, BDA, BPA | up | 1.05 | low significance |
| 5 | GlcNAc(b1-3)GalNAc | [Oglycan_core3] | AIA, UEA-II, WGA_1, WGA_2 | up | 0.86 | low significance |
| 26 | Gal(a1-3) | [] | GS-I_1, GS-I_2, GS-I_3, GS-I_4, MNA-G, PA-IL | up | 0.83 | low significance |
| 27 | Gal(a1-4) | [] | GS-I_1, GS-I_2, GS-I_3, GS-I_4, MNA-G, PA-IL | up | 0.83 | low significance |
| 30 | Gal(b1-4)GlcNAc(b1-3) | [Terminal_LacNAc_type2] | LEA_1, LEA_2, STA, BDA, BPA, ECA, RCA120, Rici... | up | 0.54 | low significance |
| 25 | Gal(a1-3)Gal | [] | EEA, EEL, MOA, GS-I_1, GS-I_2, GS-I_3, GS-I_4,... | up | 0.51 | low significance |
| 33 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc | [Internal_LacNAc_type2] | MAA_1, MAA_2, MAL-I, BDA, BPA, ECA, RCA120, Ri... | up | 0.49 | low significance |
| 37 | Gal(a1-3)GalNAc | [] | MOA, EEA, EEL, GS-I_1, GS-I_2, GS-I_3, GS-I_4,... | up | 0.46 | low significance |
| 20 | GalNAc(a1-4) | [] | GHA, HAA, HPA, CSA, GS-I_1, GS-I_2, GS-I_3, GS... | up | 0.39 | low significance |
| 19 | GalNAc(a1-3) | [] | GHA, HAA, HPA, CSA, GS-I_1, GS-I_2, GS-I_3, GS... | up | 0.39 | low significance |
| 21 | GalNAc(a1-3)GalNAc(b1-3) | [] | DBA, SBA, CSA, GHA, HAA, HPA, VVA_1, VVA_2, WF... | up | 0.25 | low significance |
| 24 | GalNAc(b1-4)GlcNAc | [Terminal_LacdiNAc_type2] | ECA, STA, CSA, SBA, VVA_1, VVA_2, WFA, BPA, WG... | up | 0.20 | low significance |
| 44 | Fuc(a1-2)Gal(b1-4)GalNAc(b1-3) | [] | SNA-II, AAA, AAL, AOL, BDA, BPA, CSA, SBA, VVA... | up | 0.16 | low significance |
| 13 | GalNAc(b1-4) | [] | CSA, SBA, VVA_1, VVA_2, WFA, BPA, WGA_1, WGA_2 | up | 0.13 | low significance |
| 12 | GalNAc(b1-3) | [] | CSA, SBA, VVA_1, VVA_2, WFA, BPA, WGA_1, WGA_2 | up | 0.13 | low significance |
| 40 | Fuc(a1-2)Gal(b1-4)GlcNAc | [H_antigen_type2, Internal_LacNAc_type2] | PTL-II, TJA-II, UEA-I, UEA-II, AAA, AAL, AOL, ... | up | 0.13 | low significance |
| 28 | GlcNAc(a1-3) | [] | HAA, HPA, WGA_1, WGA_2 | up | 0.12 | low significance |
| 29 | GlcNAc(a1-4) | [] | HAA, HPA, WGA_1, WGA_2 | up | 0.12 | low significance |
| 32 | Gal3S(b1-4)GlcNAc | [] | MAA_1, MAA_2, MAL-I, MAL-II | down | 0.12 | low significance |
| 0 | Fuc(a1-2) | [] | AAA, AAL, AOL | up | 0.09 | low significance |
| 36 | Gal3S(b1-4) | [] | MAL-II | down | 0.08 | low significance |
| 35 | Gal3S(b1-3) | [] | MAL-II | down | 0.08 | low significance |
| 49 | Fuc(a1-2)Gal(b1-4)GalNAc | [] | UEA-II, AAA, AAL, AOL, BDA, BPA | up | 0.07 | low significance |
| 9 | Gal(b1-4) | [] | BDA, BPA | up | 0.05 | low significance |
| 8 | Gal(b1-3) | [] | BDA, BPA | up | 0.05 | low significance |
| 1 | Fuc(a1-3) | [] | AAL, AOL, Lotus | down | 0.03 | low significance |
| 2 | Fuc(a1-4) | [] | AAL, AOL | down | 0.03 | low significance |
| 31 | GlcNAc(b1-4)GlcNAc(b1-4) | [Chitobiose] | LEA_1, LEA_2, WGA_1, WGA_2 | down | 0.01 | low significance |
| 50 | GlcNAc(b1-3) | [] | WGA_1, WGA_2 | down | 0.01 | low significance |
| 51 | GlcNAc(b1-4) | [] | WGA_1, WGA_2 | down | 0.01 | low significance |
| 45 | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4) | [Chitobiose] | STA, LEA_1, LEA_2, WGA_1, WGA_2 | down | 0.00 | low significance |
| 48 | GlcNAc(b1-3)Gal | [] | UEA-II, WGA_1, WGA_2 | up | 0.00 | low significance |
| 52 | Neu5Ac(a2-3) | [] | WGA_1, WGA_2 | down | 0.00 | low significance |
| 53 | Neu5Ac(a2-6) | [] | WGA_1, WGA_2 | down | 0.00 | low significance |
| 54 | Neu5Ac(a2-8) | [] | WGA_1, WGA_2 | down | 0.00 | low significance |
def get_glycoshift_per_site(
df:pandas.DataFrame | str | pathlib.Path, # DataFrame with rows formatted as 'protein_site_composition' in col 1, abundances in remaining cols
group1:list, # First group indices/names or group labels for multi-group
group2:list, # Second group indices/names
paired:bool=False, # Whether samples are paired
impute:bool=True, # Replace zeros with Random Forest model
min_samples:float=0.2, # Min percent of non-zero samples required
gamma:float=0.1, # Uncertainty parameter for CLR transform
custom_scale:float | dict=0, # Ratio of total signal in group2/group1 for an informed scale model (or group_idx: mean(group)/min(mean(groups)) signal dict for multivariate)
random_state:int | numpy.random._generator.Generator | None=None, # optional random state for reproducibility
)->DataFrame: # DataFrame with GLM coefficients and FDR-corrected p-values
Analyzes site-specific glycosylation changes in glycoproteomics data using generalized linear models (GLM) with compositional data normalization
You're working with an alpha of 0.07862467893233027 that has been adjusted for your sample size of 6.| Condition_coefficient | Condition_corr_pval | Condition_significant | Neu5Ac_Condition_coefficient | Neu5Ac_Condition_corr_pval | Neu5Ac_Condition_significant | complex_Condition_coefficient | complex_Condition_corr_pval | complex_Condition_significant | hybrid_Condition_coefficient | ... | dHex_Condition_significant | high_Man_Condition_coefficient | high_Man_Condition_corr_pval | high_Man_Condition_significant | HexNAc_Condition_coefficient | HexNAc_Condition_corr_pval | HexNAc_Condition_significant | Hex_Condition_coefficient | Hex_Condition_corr_pval | Hex_Condition_significant | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| sp|P47710|CASA1_69 | 0.353257 | 0.000000e+00 | True | 0.353257 | 0.000000e+00 | True | 0.000000 | 1.000000e+00 | False | 0.353257 | ... | True | 0.000000 | 1.000000 | False | 1.413027 | 0.000000e+00 | True | -1.563416 | 0.000000e+00 | True |
| sp|P01024|CO3_85 | -13.530653 | 0.000000e+00 | True | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | -13.530653 | ... | False | -13.530653 | 0.000000 | True | -27.061306 | 0.000000e+00 | True | 12.636985 | 0.000000e+00 | True |
| sp|P10909|CLUS_103 | -0.149909 | 0.000000e+00 | True | 4.306665 | 0.000000e+00 | True | 4.456574 | 0.000000e+00 | True | -4.606483 | ... | True | 0.000000 | 1.000000 | False | -0.599635 | 0.000000e+00 | True | -0.749544 | 0.000000e+00 | True |
| sp|Q13410|BT1A1_55 | -13.100965 | 1.331366e-72 | True | -17.275380 | 3.955343e-111 | True | -4.174415 | 7.070812e-177 | True | -8.926549 | ... | False | 0.000000 | 1.000000 | False | 12.462657 | 2.362957e-76 | True | -0.638308 | 6.203369e-16 | True |
| sp|P01011|AACT_106 | -0.027608 | 3.078766e-16 | True | -2.620928 | 0.000000e+00 | True | -2.593321 | 0.000000e+00 | True | 2.565713 | ... | True | 0.000000 | 1.000000 | False | -0.110431 | 3.078766e-16 | True | -0.138038 | 3.848458e-16 | True |
| sp|P00709|LALBA_90 | -1.220055 | 1.372347e-07 | True | -1.773743 | 2.017993e-05 | True | -0.553687 | 6.915606e-01 | False | -0.666368 | ... | True | 0.000000 | 1.000000 | False | -4.880222 | 1.372347e-07 | True | 3.511094 | 9.427322e-06 | True |
| sp|P08571|CD14_151 | 0.002309 | 3.336465e-04 | True | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | 0.002309 | ... | False | 0.002309 | 0.001437 | True | 0.004617 | 3.336465e-04 | True | 0.013851 | 3.336465e-04 | True |
| sp|P07602|SAP_426 | 0.002851 | 6.752159e-03 | True | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | 0.002851 | ... | False | 0.000000 | 1.000000 | False | 0.005702 | 6.752159e-03 | True | 0.014255 | 6.752159e-03 | True |
| sp|P07602|SAP_101 | -0.001653 | 1.856037e-01 | False | -0.001653 | 1.798929e-01 | False | 0.000000 | 1.000000e+00 | False | -0.001653 | ... | False | 0.000000 | 1.000000 | False | -0.006613 | 1.670434e-01 | False | -0.008267 | 1.856037e-01 | False |
| sp|P07602|SAP_215 | -0.002911 | 2.018278e-01 | False | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | -0.005822 | 1.834798e-01 | False | -0.005822 | 2.018278e-01 | False |
| sp|Q08431|MFGM_238 | 0.144987 | 3.025292e-01 | False | -0.252363 | 3.799644e-01 | False | 0.000000 | 1.000000e+00 | False | 0.144987 | ... | False | -0.254360 | 0.941176 | False | 0.037612 | 5.947780e-01 | False | -0.036146 | 3.833506e-01 | False |
| sp|P25311|ZA2G_109 | 0.007570 | 3.250433e-01 | False | -0.227406 | 1.091050e-01 | False | -0.234976 | 1.699985e-01 | False | 0.242546 | ... | False | 0.000000 | 1.000000 | False | 0.030279 | 3.082506e-01 | False | 0.037849 | 2.863536e-01 | False |
| sp|P10909|CLUS_291 | 0.001798 | 3.553860e-01 | False | 0.001798 | 3.799644e-01 | False | 0.000000 | 1.000000e+00 | False | 0.001798 | ... | False | 0.000000 | 1.000000 | False | 0.005395 | 3.300013e-01 | False | 0.008992 | 2.887511e-01 | False |
| sp|P10909|CLUS_86 | -0.000590 | 4.353640e-01 | False | -0.000590 | 4.375966e-01 | False | 0.000000 | 1.000000e+00 | False | -0.000590 | ... | False | 0.000000 | 1.000000 | False | -0.002359 | 3.809435e-01 | False | -0.002949 | 3.585351e-01 | False |
| sp|Q08380|LG3BP_125 | -0.001393 | 5.111342e-01 | False | -0.001393 | 5.160489e-01 | False | 0.000000 | 1.000000e+00 | False | -0.001393 | ... | False | 0.000000 | 1.000000 | False | -0.005570 | 4.510007e-01 | False | -0.006963 | 3.833506e-01 | False |
| sp|P0C0L5|CO4B_HUMAN/sp|P0C0L4|CO4A | 0.000775 | 5.322439e-01 | False | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | 0.000775 | ... | False | 0.000775 | 0.941176 | False | 0.001550 | 4.731057e-01 | False | 0.006973 | 4.055191e-01 | False |
| sp|P10909|CLUS_374 | -0.001409 | 5.699737e-01 | False | -0.001409 | 6.709711e-01 | False | 0.000000 | 1.000000e+00 | False | -0.001409 | ... | False | 0.000000 | 1.000000 | False | -0.005638 | 5.285468e-01 | False | -0.007047 | 5.111285e-01 | False |
| sp|P02788|TRFL_156 | -3.489318 | 5.699737e-01 | False | 0.698868 | 7.657590e-01 | False | -0.484459 | 8.650370e-01 | False | -3.004858 | ... | True | 0.000000 | 1.000000 | False | -4.184906 | 1.192373e-01 | False | 2.754223 | 2.863536e-01 | False |
| sp|P01833|PIGR_186 | -0.003925 | 5.699737e-01 | False | -0.040250 | 6.709711e-01 | False | -0.036325 | 8.529412e-01 | False | 0.032400 | ... | False | 0.000000 | 1.000000 | False | -0.015701 | 5.285468e-01 | False | 0.012773 | 6.584205e-01 | False |
| sp|P01871|IGHM_46 | -0.001251 | 5.699737e-01 | False | -0.001251 | 6.709711e-01 | False | 0.000000 | 1.000000e+00 | False | -0.001251 | ... | False | 0.000000 | 1.000000 | False | -0.005002 | 5.285468e-01 | False | -0.006253 | 5.059774e-01 | False |
| sp|P02788|TRFL_497 | 0.298762 | 5.699737e-01 | False | -1.945478 | 3.755496e-01 | False | -11.524302 | 2.311371e-04 | True | -13.834315 | ... | False | 0.000000 | 1.000000 | False | 1.195049 | 5.285468e-01 | False | 1.832253 | 2.887511e-01 | False |
| sp|P02749|APOH_253 | 0.000764 | 5.765965e-01 | False | 0.001528 | 6.709711e-01 | False | 0.000764 | 8.529412e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | 0.003055 | 5.285468e-01 | False | 0.003819 | 5.285468e-01 | False |
| sp|P01876|IGHA1_340 | 4.562124 | 6.627967e-01 | False | 4.150781 | 5.641617e-02 | True | -4.646176 | 8.529412e-01 | False | -2.153690 | ... | False | 3.337078 | 0.941176 | False | -0.276113 | 7.563851e-01 | False | -0.665575 | 6.974410e-01 | False |
| sp|P0DOX2|IGA2_HUMAN/sp|P01877|IGHA2 | -2.201206 | 6.628949e-01 | False | -2.015248 | 1.078886e-01 | False | -5.152422 | 2.302813e-01 | False | -3.155824 | ... | False | -4.195685 | 0.941176 | False | -0.303173 | 5.945689e-01 | False | 1.493354 | 2.863536e-01 | False |
| sp|P06858|LIPL_70 | -0.000852 | 6.628949e-01 | False | -0.001704 | 7.657590e-01 | False | -0.000852 | 8.529412e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | -0.003409 | 6.290163e-01 | False | -0.004261 | 6.290163e-01 | False |
| sp|P02790|HEMO_453 | -0.000284 | 6.937840e-01 | False | -0.000568 | 8.094146e-01 | False | -0.000284 | 8.529412e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | -0.001136 | 6.937840e-01 | False | -0.001420 | 6.937840e-01 | False |
| sp|P19652|A1AG2_HUMAN/sp|P02763|A1AG1 | -0.000680 | 7.292625e-01 | False | -0.001360 | 8.167433e-01 | False | -0.000680 | 8.529412e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | -0.002719 | 7.292625e-01 | False | -0.003399 | 7.292625e-01 | False |
| sp|P01877|IGHA2_327 | -0.692246 | 7.511404e-01 | False | 0.000000 | 1.000000e+00 | False | 0.000000 | 1.000000e+00 | False | -0.692246 | ... | False | 3.982010 | 0.941176 | False | -1.384492 | 7.511404e-01 | False | 0.179438 | 9.432450e-01 | False |
| sp|P02765|FETUA_156 | -0.000528 | 7.584045e-01 | False | -0.001057 | 8.167433e-01 | False | -0.000528 | 8.529412e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | -0.002113 | 7.584045e-01 | False | -0.002642 | 7.584045e-01 | False |
| sp|P01833|PIGR_499 | -2.996462 | 7.877926e-01 | False | -3.201522 | 2.545075e-01 | False | -1.686467 | 8.529412e-01 | False | -2.734399 | ... | False | 0.000000 | 1.000000 | False | 2.897029 | 3.082506e-01 | False | -0.906962 | 6.401825e-01 | False |
| sp|P01591|IGJ_71 | 1.352444 | 7.903746e-01 | False | 0.215802 | 8.198211e-01 | False | -0.144731 | 9.420876e-01 | False | 1.120894 | ... | False | 0.000000 | 1.000000 | False | 0.425201 | 5.285468e-01 | False | -0.931993 | 3.833506e-01 | False |
| sp|P01833|PIGR_421 | 0.337457 | 8.662386e-01 | False | 0.487402 | 6.709711e-01 | False | 0.000000 | 1.000000e+00 | False | 0.337457 | ... | False | 0.000000 | 1.000000 | False | -0.236225 | 8.245977e-01 | False | 0.026444 | 9.778702e-01 | False |
| sp|P01833|PIGR_469 | 0.914459 | 8.709747e-01 | False | -4.184570 | 3.942310e-02 | True | 14.610788 | 1.012432e-04 | True | 6.866686 | ... | True | 0.000000 | 1.000000 | False | 2.097292 | 3.427967e-01 | False | -2.417376 | 2.863536e-01 | False |
| sp|P00738|HPT_241 | 0.000030 | 9.880762e-01 | False | 0.000061 | 9.880762e-01 | False | 0.000030 | 9.880762e-01 | False | 0.000000 | ... | False | 0.000000 | 1.000000 | False | 0.000121 | 9.880762e-01 | False | 0.000152 | 9.880762e-01 | False |
34 rows × 27 columns
extract curated motifs, graph features, and sequence features from glycan sequences
def annotate_glycan(
glycan:str | networkx.classes.digraph.DiGraph, # IUPAC-condensed glycan sequence or NetworkX graph
motifs:pandas.DataFrame | None=None, # Motif dataframe (name + sequence); defaults to motif_list
termini_list:list=[], # Monosaccharide positions: 'terminal', 'internal', or 'flexible'
gmotifs:list[networkx.classes.digraph.DiGraph] | None=None, # Precalculated motif graphs for speed
)->DataFrame: # DataFrame with motif counts for the glycan
Counts occurrences of known motifs in a glycan structure using subgraph isomorphism
| motif_name | Terminal_LewisX | Internal_LewisX | LewisY | SialylLewisX | SulfoSialylLewisX | Terminal_LewisA | Internal_LewisA | LewisB | SialylLewisA | SulfoLewisA | ... | Mucin_elongated_core2 | Fucoidan | Alginate | FG | XX | Difucosylated_core | GalFuc_core | DisialylLewisC | RM2 | DisialylLewisA |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
1 rows × 165 columns
def annotate_dataset(
glycans:list, # List of IUPAC-condensed glycan sequences
motifs:pandas.DataFrame | None=None, # Motif dataframe (name + sequence); defaults to motif_list
feature_set:list=['known'], # Feature types to analyze: known, graph, exhaustive, terminal(1-3), custom, chemical, size_branch
termini_list:list=[], # Monosaccharide positions: 'terminal', 'internal', or 'flexible'
condense:bool=False, # Remove columns with only zeros
custom_motifs:list=[], # Custom motifs when using 'custom' feature set
)->DataFrame: # DataFrame mapping glycans to presence/absence of motifs
Comprehensive glycan annotation combining multiple feature types: structural motifs, graph properties, terminal sequences
glycans = ['Man(a1-3)[Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc',
'Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
'GalNAc(a1-4)GlcNAcA(a1-4)[GlcN(b1-7)]Kdo(a2-5)[Kdo(a2-4)]Kdo(a2-6)GlcN4P(b1-6)GlcN4P']
print("Annotate Test")
out = annotate_dataset(glycans)Annotate Test| motif_name | Terminal_LewisX | Internal_LewisX | LewisY | SialylLewisX | SulfoSialylLewisX | Terminal_LewisA | Internal_LewisA | LewisB | SialylLewisA | SulfoLewisA | H_antigen_type2 | H_antigen_type1 | H_antigen_type3 | A_antigen | A_antigen_type1 | A_antigen_type2 | A_antigen_type3 | B_antigen | B_antigen_type1 | B_antigen_type2 | ExtB | Galili_antigen | GloboH | Gb5 | Gb4 | Gb3 | 3SGb3 | 8DSGb3 | 3SGb4 | 8DSGb4 | 6DSGb4 | 3SGb5 | 8DSGb5 | 6DSGb5 | 6DSGb5_2 | 6SGb3 | 8DSGb3_2 | 6SGb4 | 8DSGb4_2 | 6SGb5 | 8DSGb5_2 | 66DSGb5 | Forssman_antigen | iGb3 | I_antigen | i_antigen | PI_antigen | Chitobiose | Trimannosylcore | Internal_LacNAc_type1 | Terminal_LacNAc_type1 | Internal_LacNAc_type2 | Terminal_LacNAc_type2 | Internal_LacdiNAc_type1 | Terminal_LacdiNAc_type1 | Internal_LacdiNAc_type2 | Terminal_LacdiNAc_type2 | bisectingGlcNAc | VIM | PolyLacNAc | Ganglio_Series | Lacto_Series(LewisC) | NeoLacto_Series | betaGlucan | KeratanSulfate | Hyaluronan | Mollu_series | Arthro_series | Cellulose_like | Chondroitin_4S | GPI_anchor | Isoglobo_series | LewisD | Globo_series | Sda | SDA | Muco_series | Heparin | Peptidoglycan | Dermatansulfate | CAD | Lactosylceramide | Lactotriaosylceramide | LexLex | GM3 | H_type3 | GM2 | GM1 | cisGM1 | VIM2 | GD3 | GD1a | GD2 | GD1b | SDLex | Fuc_LN3 | GT1b | GD1 | GD1a_2 | LcGg4 | GT3 | Disialyl_T_antigen | GT1a | GT2 | GT1c | 2Fuc_GM1 | GQ1c | O_linked_mannose | GT1aa | GQ1b | HNK1 | GQ1ba | O_mannose_Lex | 2Fuc_GD1b | Sialopentaosylceramide | Sulfogangliotetraosylceramide | B-GM1 | GQ1aa | bisSulfo-Lewis x | para-Forssman | core_fucose | core_fucose(a1-3) | GP1c | B-GD1b | GP1ca | Isoglobotetraosylceramide | polySia | high_mannose | Gala_series | LPS_core | Nglycan_complex | Nglycan_complex2 | Oglycan_core1 | Oglycan_core2 | Oglycan_core3 | Oglycan_core4 | Oglycan_core5 | Oglycan_core6 | Oglycan_core7 | Xylogalacturonan | Sialosylparagloboside | LDNF | OFuc | Arabinogalactan_type2 | EGF_repeat | Nglycan_hybrid | Arabinan | Xyloglucan | Acharan_Sulfate | M3FX | M3X | 1-6betaGalactan | Arabinogalactan_type1 | Galactomannan | Tetraantennary_Nglycan | Mucin_elongated_core2 | Fucoidan | Alginate | FG | XX | Difucosylated_core | GalFuc_core | DisialylLewisC | RM2 | DisialylLewisA |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Man(a1-3)[Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| GalNAc(a1-4)GlcNAcA(a1-4)[GlcN(b1-7)]Kdo(a2-5)[Kdo(a2-4)]Kdo(a2-6)GlcN4P(b1-6)GlcN4P | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
def quantify_motifs(
df:str | pandas.DataFrame, # DataFrame or filepath with samples as columns, abundances as values
glycans:list, # List of IUPAC-condensed glycan sequences
feature_set:list, # Feature types to analyze: known, graph, exhaustive, terminal(1-3), custom, chemical, size_branch
custom_motifs:list=[], # Custom motifs when using 'custom' feature set
remove_redundant:bool=True, # Remove redundant motifs via deduplicate_motifs
)->DataFrame: # DataFrame with motif abundances (motifs as columns, samples as rows)
Extracts and quantifies motif abundances from glycan abundance data by weighting motif occurrences
| control_1 | tumor_1 | control_2 | tumor_2 | control_3 | tumor_3 | control_4 | tumor_4 | control_5 | tumor_5 | ... | control_16 | tumor_16 | control_17 | tumor_17 | control_18 | tumor_18 | control_19 | tumor_19 | control_20 | tumor_20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| H_antigen_type2 | 1.347737 | 0.892651 | 2.468405 | 1.810795 | 1.589162 | 0.449339 | 2.640132 | 0.572828 | 2.763890 | 0.737076 | ... | 1.070249 | 0.647786 | 1.440912 | 1.810304 | 1.722289 | 1.475260 | 4.847788 | 4.552496 | 0.480035 | 0.494123 |
| Internal_LacNAc_type2 | 8.845085 | 10.063160 | 13.435501 | 28.834006 | 5.585973 | 11.359659 | 11.672584 | 21.193308 | 12.734919 | 28.597709 | ... | 10.883437 | 17.991155 | 21.166792 | 16.161351 | 11.909325 | 29.924308 | 12.820872 | 19.107379 | 8.802443 | 10.268911 |
| Terminal_LacNAc_type2 | 52.982192 | 13.183951 | 24.413523 | 12.870782 | 9.555884 | 9.822266 | 12.628910 | 13.916662 | 26.569737 | 10.733867 | ... | 18.779972 | 12.157928 | 14.828507 | 20.879287 | 27.689619 | 10.734756 | 28.328965 | 37.870847 | 14.835019 | 8.910804 |
| Disialyl_T_antigen | 20.803836 | 36.895471 | 32.803297 | 20.401157 | 33.971366 | 30.150599 | 37.703636 | 24.728411 | 31.798990 | 15.989214 | ... | 46.337629 | 39.476930 | 39.087708 | 40.348217 | 35.791797 | 22.968160 | 11.026029 | 2.613718 | 44.676379 | 46.125360 |
| Oglycan_core1 | 37.329013 | 75.567842 | 59.998893 | 57.608119 | 83.293693 | 78.436161 | 73.308916 | 64.356888 | 58.197862 | 60.329536 | ... | 68.269613 | 68.762287 | 62.541874 | 60.699726 | 58.713271 | 58.203265 | 58.826129 | 42.904325 | 74.390026 | 79.515568 |
| Mucin_elongated_core2 | 61.827277 | 23.247111 | 37.849024 | 41.704788 | 15.141858 | 21.181925 | 24.301494 | 35.109970 | 39.304656 | 39.331576 | ... | 29.663409 | 30.149083 | 35.995300 | 37.040638 | 39.598944 | 40.659064 | 41.149838 | 56.978227 | 23.637462 | 19.179715 |
| Gal | 163.691481 | 126.500106 | 141.895063 | 147.702533 | 115.056369 | 132.721945 | 122.804259 | 138.398297 | 141.412183 | 167.203077 | ... | 133.838024 | 140.218313 | 142.530133 | 139.697255 | 138.848449 | 154.791018 | 142.588964 | 157.426027 | 122.916027 | 120.555251 |
| GalNAc | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | ... | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 | 100.000000 |
| GalOS | 0.843710 | 1.185047 | 2.152084 | 0.687093 | 1.564450 | 0.381914 | 2.389590 | 0.533142 | 2.497482 | 0.338889 | ... | 2.066978 | 1.088630 | 1.462826 | 2.259636 | 1.687785 | 1.137672 | 0.024033 | 0.117449 | 1.972512 | 1.304717 |
| GlcNAc6S(b1-6)GalNAc | 2.707913 | 4.438043 | 6.198123 | 6.684838 | 1.478960 | 11.921934 | 0.892356 | 3.821469 | 4.605009 | 28.210391 | ... | 6.241593 | 11.157860 | 7.997660 | 4.916252 | 0.937290 | 15.269626 | 1.463159 | 0.565249 | 1.251077 | 2.680253 |
| Neu5Ac | 80.494155 | 134.094482 | 120.708503 | 125.892731 | 128.626161 | 137.543517 | 132.135127 | 124.740497 | 118.279272 | 134.227059 | ... | 149.089683 | 152.360772 | 145.124475 | 140.251427 | 125.331418 | 121.962226 | 91.599064 | 72.000898 | 142.956534 | 148.579697 |
| Neu5Ac(a2-3)Gal | 57.345927 | 94.670033 | 83.675402 | 103.574200 | 91.775344 | 106.231617 | 90.136699 | 98.461821 | 81.110136 | 117.087919 | ... | 97.928245 | 109.749014 | 101.760261 | 93.222423 | 86.403840 | 96.715461 | 80.029183 | 69.040921 | 95.565848 | 99.973512 |
| Neu5Ac(a2-6)GalNAc | 22.219773 | 38.119351 | 34.492798 | 21.576036 | 35.006672 | 30.847852 | 39.239054 | 25.722979 | 34.591496 | 16.715338 | ... | 48.608621 | 40.893926 | 41.366216 | 44.132349 | 36.994779 | 23.981142 | 11.515995 | 2.819886 | 45.199008 | 46.969521 |
| Neu5Ac(a2-8)Neu5Ac | 0.084745 | 0.120050 | 0.388219 | 0.055402 | 0.279696 | 0.082135 | 0.369784 | 0.022555 | 0.080158 | 0.084913 | ... | 0.485839 | 0.629202 | 0.535171 | 0.637019 | 0.245015 | 0.127952 | 0.029853 | 0.022643 | 0.219166 | 0.331947 |
| Gal(b1-3)GalNAc | 99.156290 | 98.814953 | 97.847916 | 99.312907 | 98.435550 | 99.618086 | 97.610410 | 99.466858 | 97.502518 | 99.661111 | ... | 97.933022 | 98.911370 | 98.537174 | 97.740364 | 98.312215 | 98.862328 | 99.975967 | 99.882551 | 98.027488 | 98.695283 |
15 rows × 40 columns
def get_k_saccharides(
glycans:list[str] | set[str], # List or set of IUPAC-condensed glycan sequences
size:int=2, # Number of monosaccharides per fragment
up_to:bool=False, # Include fragments up to size k (adds monosaccharides)
just_motifs:bool=False, # Return nested list of motifs instead of count DataFrame
terminal:bool=False, # Only count terminal fragments
)->pandas.DataFrame | list[list[str]]: # DataFrame of k-saccharide counts or list of motifs per glycan
Extracts k-saccharide fragments from glycan sequences with options for different fragment sizes and positions
| Kdo(a2-4)Kdo | Kdo(a2-4/5)Kdo | Man(a1-2)Man | Man(a1-6)Man | Fuc(a1-3)GlcNAc | Man(a1-3)Man | Man(b1-4)GlcNAc | GlcNAcA(a1-4)Kdo | Man(a1-2/3/6)Man | Kdo(a2-5)Kdo | GalNAc(a1-4)GlcNAcA | Xyl(b1-2)Man | GlcN(b1-7)Kdo | Kdo(a2-6)GlcN4P | GlcNAc(b1-4)GlcNAc | GlcN4P(b1-6)GlcN4P | Xyl(b1-2)Man(b1-4)GlcNAc | Kdo(a2-5)Kdo(a2-6)GlcN4P | Kdo(a2-4)[Kdo(a2-5)]Kdo | Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAcA(a1-4)[GlcN(b1-7)]Kdo | Kdo(a2-4)Kdo(a2-6)GlcN4P | Xyl(b1-2)[Man(a1-3/6)]Man | Fuc(a1-3)[GlcNAc(b1-4)]GlcNAc | GalNAc(a1-4)GlcNAcA(a1-4)Kdo | Man(a1-3/6)Man(b1-4)GlcNAc | Kdo(a2-4/5)Kdo(a2-6)GlcN4P | Man(a1-2/3)Man(a1-2/3/6)Man | Man(a1-2)Man(a1-3)Man | Man(a1-6)Man(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc | Kdo(a2-6)GlcN4P(b1-6)GlcN4P | Xyl(b1-2)[Man(a1-6)]Man | GlcN(b1-7)Kdo(a2-5)Kdo | Man(a1-3)[Man(a1-6)]Man | GlcNAcA(a1-4)Kdo(a2-5)Kdo | Man(a1-3)Man(a1-6)Man | Man(a1-2)Man(a1-2)Man | Xyl(b1-2)[Man(a1-3)]Man | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 2 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 2 | 1 | 0 | 2 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 1 |
| 1 | 0 | 0 | 2 | 1 | 0 | 2 | 1 | 0 | 5 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 3 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 |
| 2 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 2 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 0 |
Identifies terminal monosaccharide sequences from non-reducing ends of glycan structure
['Neu5Ac(a2-3)', 'Neu5Ac(a2-6)']Retrieves molecular properties from PubChem for a list of glycans using their SMILES representations
| undefined_atom_stereo_count | isotope_atom_count | defined_atom_stereo_count | bond_stereo_count | undefined_bond_stereo_count | defined_bond_stereo_count | tpsa | monoisotopic_mass | xlogp | molecular_weight | atom_stereo_count | complexity | h_bond_donor_count | heavy_atom_count | exact_mass | h_bond_acceptor_count | covalent_unit_count | charge | rotatable_bond_count | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | 1 | 0 | 56 | 0 | 0 | 0 | 1070 | 2222.7830048 | -23.600000 | 2224.0 | 57 | 4410 | 39 | 152 | 2222.7830048 | 62 | 1 | 0 | 43 |
def get_glycan_similarity(
glycan1:str | networkx.classes.digraph.DiGraph, # IUPAC-condensed glycan sequence or NetworkX graph
glycan2:str | networkx.classes.digraph.DiGraph, # IUPAC-condensed glycan sequence or NetworkX graph
motifs:pandas.DataFrame | None=None, # Motif dataframe (name + sequence); defaults to motif_list
feature_set:list=['known', 'exhaustive', 'terminal'], # Feature types to analyze: known, graph, exhaustive, terminal(1-3), custom, chemical, size_branch
)->float: # Cosine similarity between glycan1 and glycan2
Calculates cosine similarity between two glycans based on their motif count fingerprints
convert glycan sequences to graphs and contains helper functions to search for motifs / check whether two sequences describe the same sequence, etc.
def glycan_to_nxGraph(
glycan:str, # Glycan in IUPAC-condensed format
libr:glycowork.glycan_data.loader.HashableDict[str, int] | None=None, # Dictionary of form glycoletter:index
termini:str='ignore', # How to encode terminal/internal position; options: ignore, calc, provided
termini_list:tuple[str] | None=None, # List of positions from terminal/internal/flexible
)->DiGraph: # NetworkX graph object of glycan
Wrapper for converting glycans into networkx graphs; also works with floating substituents
Glycan to networkx Graph (only edges printed)
[(1, 0), (3, 2), (4, 1), (4, 3), (5, 4), (6, 5), (7, 6), (9, 8), (10, 7), (10, 9)]Convert glycan graph back to IUPAC-condensed format, handling disconnected components
'Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc'Check whether two glycans are identical
Graph Isomorphism Test
True
def subgraph_isomorphism(
glycan:str | networkx.classes.digraph.DiGraph, # Glycan sequence or graph
motif:str | networkx.classes.digraph.DiGraph, # Glycan motif sequence or graph
termini_list:list=[], # List of monosaccharide positions from terminal/internal/flexible
count:bool=False, # Whether to return count instead of presence/absence
return_matches:bool=False, # Whether to return matched subgraphs as node lists
)->bool | int | tuple[int, list[list[int]]]: # Boolean presence, count, or (count, matches)
Check if motif exists as subgraph in glycan
Subgraph Isomorphism Test
TrueCompute graph features of glycan or network
| diameter | branching | nbrLeaves | avgDeg | varDeg | maxDeg | nbrDeg4 | max_deg_leaves | mean_deg_leaves | deg_assort | ... | flow_edgeMax | flow_edgeMin | flow_edgeAvg | flow_edgeVar | secorderMax | secorderMin | secorderAvg | secorderVar | egap | entropyStation | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | 8 | 1 | 3 | 1.818182 | 0.330579 | 3.0 | 0 | 3.0 | 3.0 | -1.850372e-15 | ... | 0.333333 | 0.111111 | 0.217778 | 0.007289 | 45.607017 | 20.736441 | 31.679285 | 62.422895 | 0.340654 | -2.180184 |
1 rows × 49 columns
Find the largest common subgraph of two glycans
'Fuc(a1-6)GlcNAc'Ensures function compatibility with string glycans and graph glycans
<networkx.classes.digraph.DiGraph>
def get_possible_topologies(
glycan:str | networkx.classes.digraph.DiGraph, # Glycan with floating substituent
exhaustive:bool=False, # Whether to allow additions at internal positions
allowed_disaccharides:set[str] | None=None, # Permitted disaccharides when creating possible glycans
modification_map:dict={'6S': {'Gal', 'GlcNAc'}, '3S': {'Gal'}, '4S': {'GalNAc'}, 'OS': {'Gal', 'GalNAc', 'GlcNAc'}}, # Maps modifications to valid attachments
return_graphs:bool=False, # Whether to return glycan graphs (otherwise return converted strings)
)->list: # List of possible topology strings or graphs
Create possible glycan graphs given a floating substituent
Check whether glycan with floating substituent could match glycans from a list
['Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc']Remove duplicate glycans from a list/set, even if they have different strings
['Neu5Ac(a2-6)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc',
'Fuc(a1-2)Gal(b1-3)GalNAc',
'Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)Gal(b1-3)]GalNAc']process IUPAC-condensed glycan sequences into glycoletters etc.
Convert list of glycans into a nested lists of glycoletters
[['Man', 'a1-3', 'Man', 'a1-6', 'Man', 'b1-4', 'GlcNAc', 'b1-4', 'GlcNAc'],
['Man',
'a1-2',
'Man',
'a1-3',
'Man',
'a1-6',
'Man',
'b1-4',
'GlcNAc',
'b1-4',
'GlcNAc']]Returns dictionary mapping glycoletters to indices
{'GlcNAc': 0, 'Man': 1, 'a1-2': 2, 'a1-3': 3, 'a1-6': 4, 'b1-4': 5}Updates libr with newly introduced glycoletters
{'GlcNAc': 0, 'Man': 1, 'a1-2': 2, 'a1-3': 3, 'a1-6': 4, 'b1-4': 5, 'Fuc': 6}Converts a dataframe with glycan occurrence to absence/presence matrix
| glycan | Apif(a1-2)Xyl(b1-2)[Glc6Ac(b1-4)]Glc | Ara(a1-2)Ara(a1-6)GlcNAc | Ara(a1-2)Glc(b1-2)Ara | Ara(a1-2)GlcA | Ara(a1-2)[Glc(b1-6)]Glc | Ara(a1-3)Gal(b1-6)Gal | Ara(a1-6)Glc | Araf(a1-3)Araf(a1-5)[Araf(a1-6)Gal(b1-6)Glc(b1-6)Man(a1-3)]Araf(a1-5)Araf(a1-3)Araf(a1-3)Araf | Araf(a1-3)Gal(b1-6)Gal | D-Apif(b1-2)Glc | D-Apif(b1-2)GlcA | D-Apif(b1-3)Xyl(b1-2)[Glc6Ac(b1-4)]Glc | D-Apif(b1-3)Xyl(b1-4)Rha(a1-2)Ara | D-Apif(b1-3)Xyl(b1-4)Rha(a1-2)D-Fuc | D-Apif(b1-3)Xyl(b1-4)[Glc(b1-3)]Rha(a1-2)D-Fuc | D-Apif(b1-3)[Gal(b1-4)Xyl(b1-4)]Rha(a1-2)D-Fuc | D-Apif(b1-3)[Gal(b1-4)Xyl(b1-4)]Rha(a1-2)[Rha(a1-3)]D-Fuc | D-Apif(b1-3)[Gal(b1-4)Xyl(b1-4)]Rha(a1-3)D-Fuc | D-Apif(b1-6)Glc | D-ApifOMe(b1-3)XylOMe(b1-4)RhaOMe(a1-2)D-FucOMe | D-ApifOMe(b1-3)XylOMe(b1-4)[GlcOMe(b1-3)]RhaOMe(a1-2)D-FucOMe | Fruf(a2-1)[Glc(b1-2)][Glc(b1-3)Glc4Ac6Ac(b1-3)]Glc | Fruf(a2-1)[Glc(b1-2)][Glc(b1-3)Glc4Ac6Ac(b1-3)]Glc6Ac | Fruf(a2-1)[Glc(b1-2)][Glc(b1-3)Glc6Ac(b1-3)]Glc | Fruf(a2-1)[Glc(b1-2)][Glc(b1-3)Glc6Ac(b1-3)]Glc6Ac | Fruf(b2-1)Glc3Ac6Ac | Fruf(b2-1)Glc4Ac6Ac | Fruf(b2-1)Glc6Ac | Fruf(b2-1)[Glc(b1-2)]Glc | Fruf(b2-1)[Glc(b1-2)][Glc(b1-3)Glc(b1-3)]Glc | Fruf(b2-1)[Glc(b1-2)][Glc(b1-3)]Glc6Ac | Fruf(b2-1)[Glc(b1-2)][Glc(b1-4)Glc(b1-3)]Glc | Fruf(b2-1)[Glc(b1-2)][Glc(b1-4)Glc(b1-3)]Glc6Ac | Fruf(b2-1)[Glc(b1-2)][Glc(b1-4)Glc6Ac(b1-3)]Glc | Fruf(b2-1)[Glc(b1-2)][Glc(b1-4)Glc6Ac(b1-3)]Glc6Ac | Fruf(b2-1)[Glc(b1-2)][Glc6Ac(b1-3)]Glc | Fruf(b2-1)[Glc(b1-2)][Glc6Ac(b1-3)]Glc6Ac | Fruf(b2-1)[Glc(b1-4)Glc6Ac(b1-3)]Glc6Ac | Fruf(b2-1)[Glc3Ac(b1-2)]Glc | Fruf(b2-1)[Glc6Ac(b1-2)]Glc | Fruf1Ac(b2-1)Glc2Ac4Ac6Ac | Fuc(a1-2)Gal(b1-2)Xyl(a1-6)Glc | Fuc(a1-2)Gal(b1-2)Xyl(a1-6)Glc(b1-4)Glc | Fuc(a1-2)Gal(b1-2)Xyl(a1-6)[Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)]Glc(b1-4)Glc | Fuc(a1-2)Gal(b1-2)Xyl(a1-6)[Glc(b1-4)]Glc(b1-4)Glc | Fuc(a1-2)Gal(b1-4)Xyl | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-6)GlcNAc(b1-2)[Man(a1-6)]Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(?1-?)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Man(a1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(?1-?)[Gal(?1-?)]GlcNAc(?1-?)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(?1-?)Man(a1-3)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(a1-4)Gal | Gal(a1-6)Gal | Gal(a1-6)Gal(a1-6)Gal | Gal(a1-6)Gal(a1-6)Gal(a1-6)Gal(a1-6)Glc(a1-2)Fru | Gal(a1-6)Gal(a1-6)Gal(a1-6)Gal(a1-6)Glc(a1-2)Fruf | Gal(a1-6)Gal(a1-6)Gal(a1-6)Gal(a1-6)[Fruf(b2-1)]Glc | Gal(a1-6)Gal(a1-6)Gal(a1-6)Glc | Gal(a1-6)Gal(a1-6)Gal(a1-6)Glc(a1-2)Fru | Gal(a1-6)Gal(a1-6)Gal(a1-6)Glc(a1-2)Fruf | Gal(a1-6)Gal(a1-6)Glc | Gal(a1-6)Gal(a1-6)Glc(a1-2)Fru | Gal(a1-6)Gal(a1-6)Glc(a1-2)Fruf | Gal(a1-6)Glc(a1-2)Fru | Gal(a1-6)Glc(a1-2)Fruf | Gal(a1-6)Man | Gal(a1-6)Man(b1-4)Man | Gal(a1-6)Man(b1-4)Man(b1-4)Man(b1-4)Man | Gal(a1-6)Man(b1-4)Man(b1-4)Man(b1-4)[Gal(a1-6)]Man(b1-4)Man(b1-4)Man(b1-4)[Gal(a1-6)]Man | Gal(a1-6)Man(b1-4)Man(b1-4)[Gal(a1-6)]Man | Gal(a1-6)Man(b1-4)[Gal(a1-6)]Man | Gal(b1-2)Glc | Gal(b1-2)GlcA | Gal(b1-2)GlcA6Me | Gal(b1-2)Xyl(a1-6)Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Gal(b1-2)Xyl(a1-6)[Glc(b1-4)]Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Gal(b1-2)Xyl(a1-6)[Glc(b1-4)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Gal(b1-2)[Xyl(b1-3)]GlcA | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-4)Man(a1-3)[Gal(b1-3)GlcNAc(b1-4)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-4)Man(a1-3)[GlcNAc(b1-4)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-4)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-4)Man(a1-6)[GlcNAc(b1-4)Man(a1-3)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-4)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3/6)[Gal(b1-3)GlcNAc(b1-2)Man(a1-3/6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3/6)[GlcNAc(b1-2)Man(a1-3/6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3/6)[Xyl(b1-2)][Man(a1-3/6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)[Man(a1-6)]Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-6)]GlcNAc(b1-2)[Man(a1-6)]Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-4)Gal(b1-4)Man | Gal(b1-4)Gal(b1-4)ManOMe | Gal(b1-4)GlcA | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)Man(b1-4)Man | Gal(b1-4)Man(b1-4)Man(b1-4)Gal | Gal(b1-4)Xyl(b1-4)Rha(a1-2)D-Fuc | Gal(b1-4)Xyl(b1-4)Rha(a1-2)D-Fuc1CoumOMe | Gal(b1-4)Xyl(b1-4)Rha(a1-2)D-Fuc1FerOMe | Gal(b1-4)Xyl(b1-4)Rha(a1-2)Fuc | Gal(b1-4)Xyl(b1-4)Rha(a1-2)Fuc4Ac | Gal(b1-4)Xyl(b1-4)Rha(a1-2)[Rha(a1-3)]D-Fuc | Gal(b1-4)Xyl(b1-4)Rha(a1-2)[Rha(a1-3)]D-Fuc1CoumOMe | Gal(b1-4)Xyl(b1-4)Rha(a1-2)[Rha(a1-3)]D-FucOMeOSin | Gal(b1-4)Xyl(b1-4)Rha(a1-2)[Rha(a1-3)]Fuc | Gal(b1-4)Xyl(b1-4)[D-Apif(b1-3)]Rha(a1-2)D-Fuc | Gal(b1-4)Xyl(b1-4)[D-Apif(b1-3)]Rha(a1-2)D-Fuc1CoumOMe | Gal(b1-4)Xyl(b1-4)[D-Apif(b1-3)]Rha(a1-2)[Rha(a1-3)]D-Fuc | Gal(b1-4)Xyl(b1-4)[D-Apif(b1-3)]Rha(a1-2)[Rha(a1-3)]D-Fuc1CoumOMe | GalA(a1-2)[Araf(a1-5)Araf(a1-4)]Rha(b1-4)GalA | GalA(a1-4)GalA(a1-4)GalA | GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-2)Rha(a1-4)GalA(a1-2)Rha(a1-4)GalA(a1-2)GalA | GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA(a1-4)GalA | GalOMe(b1-2)[XylOMe(b1-3)]GlcAOMe | GalOMe(b1-4)XylOMe(b1-4)RhaOMe(a1-2)D-FucOMe | GalOMe(b1-4)XylOMe(b1-4)RhaOMe(a1-2)[RhaOMe(a1-3)]D-FucOMe | GalOMe(b1-4)XylOMe(b1-4)[D-ApifOMe(b1-3)]RhaOMe(a1-2)[RhaOMe(a1-3)]D-FucOMe | Galf(b1-2)[Galf(b1-4)]Man | Glc(a1-2)Fru | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Glc(a1-2)Rha(a1-6)Glc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-4)Glc(a1-2)Rha(a1-6)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-6)Glc | Glc(a1-4)Glc(a1-4)GlcA | Glc(a1-4)GlcA(b1-2)GlcA | Glc(b1-2)Ara | Glc(b1-2)Ara(a1-2)GlcA | Glc(b1-2)Gal(b1-2)Gal(b1-2)GlcA | Glc(b1-2)Gal(b1-2)GlcA | Glc(b1-2)Gal(b1-2)GlcA(b1-3)[Glc(b1-3)]Ara | Glc(b1-2)Glc | Glc(b1-2)Glc(a1-2)Fru | Glc(b1-2)Glc(a1-2)FrufOBzOCin | Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)GlcA | Glc(b1-2)Xyl | Glc(b1-2)[Ara(a1-3)]GlcA6Me | Glc(b1-2)[Ara(a1-3)]GlcAOMe | Glc(b1-2)[Ara(a1-6)]Glc | Glc(b1-2)[Glc(b1-3)]Glc(a1-2)Fruf | Glc(b1-2)[Glc(b1-3)]Glc1Fer6Ac(a1-2)Fruf1FerOBz | Glc(b1-2)[Glc(b1-3)]Glc6Ac(a1-2)Fru | Glc(b1-2)[Glc6Ac(b1-3)]Glc(a1-2)Fru | Glc(b1-2)[Glc6Ac(b1-3)]Glc1Fer(a1-2)Fruf1FerOBz | Glc(b1-2)[Glc6Ac(b1-3)]Glc6Ac(a1-2)Fru | Glc(b1-2)[Rha(a1-3)]GlcA | Glc(b1-2)[Xyl(b1-2)Ara(a1-6)]Glc | Glc(b1-2)[Xyl(b1-2)D-Fuc(b1-6)]Glc | Glc(b1-3)Ara | Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)[Glc(b1-2)]Glc(a1-2)Fru | Glc(b1-3)Glc(b1-3)[Glc(b1-2)]Glc(a1-2)Fruf | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc(a1-2)Fru | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc(a1-2)Fruf | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Coum6Ac(a1-2)Fruf1CoumOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1CoumOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1FerOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1CoumOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1FerOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)]Glc6Ac(a1-2)Fru | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)][Rha(a1-4)]Glc1Coum6Ac(a1-2)Fruf1CoumOBz | Glc(b1-3)Glc6Ac(b1-3)[Glc(b1-2)][Rha(a1-4)]Glc1Fer6Ac(a1-2)Fruf1CoumOBz | Glc(b1-3)Rha1Fer(a1-4)Fruf(b2-1)GlcOBz | Glc(b1-3)[Araf(a1-4)]Rha(a1-2)Glc | Glc(b1-3)[Xyl(b1-4)]Rha(a1-2)D-FucOMe | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc(a1-2)Fru | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc(a1-2)Fruf | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc1Coum6Ac(a1-2)Fruf1FerOBz | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1FerOBz | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1CoumOBz | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1FerOBz | Glc(b1-4)Glc(b1-3)[Glc(b1-2)]Glc6Ac(a1-2)Fru | Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Man | Glc(b1-4)Glc6Ac(b1-3)Glc1Fer6Ac(a1-2)Fruf1FerOBz | Glc(b1-4)Glc6Ac(b1-3)Glc6Ac(a1-2)Fru | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc(a1-2)Fru | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Coum6Ac(a1-2)Fruf1FerOBz | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1FerOBz | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1CoumOBz | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1FerOBz | Glc(b1-4)Glc6Ac(b1-3)[Glc(b1-2)]Glc6Ac(a1-2)Fru | Glc(b1-4)Man(b1-4)Glc | Glc(b1-4)Rha | Glc(b1-4)Rha1Fer(a1-4)Fruf(b2-1)GlcOBz | Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Glc(b1-6)Glc(b1-3)Glc | Glc(b1-6)GlcNAc | Glc1Cer | Glc2Ac(b1-4)[D-Apif(b1-3)Xyl(b1-2)]Glc | Glc2Ac3Ac4Ac6Ac(b1-3)Ara | Glc3Ac(b1-2)Glc(a1-2)Fru | Glc6Ac(a1-2)Fru | Glc6Ac(b1-2)Glc(a1-2)Fru | Glc6Ac(b1-2)Glc(a1-2)FrufOBzOCin | Glc6Ac(b1-3)Ara | Glc6Ac(b1-3)Glc6Ac(b1-3)[Glc6Ac(b1-2)]Glc1Fer6Ac(a1-2)Fruf1CoumOAcOBz | Glc6Ac(b1-3)Glc6Ac(b1-3)[Glc6Ac(b1-2)][RhaOAc(a1-4)]Glc1Fer6Ac(a1-2)Fruf1CoumOAcOBz | Glc6Ac(b1-3)[Glc(b1-2)]Glc1Coum(a1-2)Fruf1CoumOBz | Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1CoumOBz | Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer(a1-2)Fruf1FerOBz | Glc6Ac(b1-3)[Glc(b1-2)]Glc1Fer6Ac(a1-2)Fruf1FerOBz | GlcA(b1-2)Glc | GlcA(b1-2)GlcA | GlcA(b1-2)GlcA(b1-2)Rha | GlcA4Me(a1-2)[Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)]Xyl | GlcA4Me(a1-2)[Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)]Xyl | GlcA4Me(a1-2)[Xyl(b1-4)]Xyl | GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Gal(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3/6)[Xyl(b1-2)][Man(a1-3/6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3/6)[Xyl(b1-2)][Man(a1-3/6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-4)Man(a1-3)[GlcNAc(b1-4)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-4)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-4)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcOMe(b1-3)[XylOMe(b1-4)]RhaOMe(a1-2)D-FucOMe | Glcf(b1-2)Xyl(b1-4)Rha(b1-4)[Xyl(b1-3)]Xyl | Hexf(?1-?)Xyl(b1-4)Rha(b1-4)[Xyl(a1-3)]Xyl | L-Lyx(a1-2)Ara(a1-2)GlcA | Lyx(a1-2)Ara(a1-2)GlcA | Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-6)[Man(a1-2)Man(a1-3)]Man(a1-3)[Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)[Man(a1-6)]Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-3)[Man(a1-2)Man(a1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-2)Man(a1-6)[Man(a1-2)Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAcN | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)[Man(a1-3)]Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-3)[Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-6)][Xylf(a1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc-ol | Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAcN | Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]Hex | Man(a1-3)[Xylf(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3/6)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-3/6)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3/6)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3/6)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(b1-2)Man | Man(b1-4)Gal(b1-4)Gal(b1-4)Man | Man(b1-4)Gal(b1-4)Gal(b1-4)ManOMe | Man(b1-4)Man | Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)[Gal(a1-6)]Man | Man(b1-4)Man(b1-4)[Gal(a1-6)]Man | Man(b1-4)Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)Man(b1-4)Man(b1-4)[Man(b1-6)]Man(b1-4)[Man(b1-6)]Man(b1-4)Man(b1-4)Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-3)Gal(a1-3)Gal(a1-6)]Man(b1-4)Man(b1-4)Man(b1-4)[Man(b1-6)]Man(b1-4)[Man(b1-6)]Man(b1-4)Man(b1-4)Man(b1-4)[Man(b1-6)]Man(b1-4)[Man(b1-6)]Man(b1-4)Man(b1-4)Man | Man(b1-4)[Gal(a1-6)]Man | Man(b1-4)[Gal(a1-6)]Man(b1-4)Man | Man(b1-4)[Gal(a1-6)]Man(b1-4)Man(b1-4)Man | Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man | Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)Man | Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man | Man(b1-6)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Rha(a1-2)Ara | Rha(a1-2)Ara(a1-2)GlcA | Rha(a1-2)Ara(a1-2)GlcA6Me | Rha(a1-2)Ara(a1-2)GlcAOMe | Rha(a1-2)D-Ara(b1-2)GlcA | Rha(a1-2)Gal(b1-2)Glc | Rha(a1-2)Gal(b1-2)GlcA | Rha(a1-2)Gal(b1-2)GlcA6Me | Rha(a1-2)Gal(b1-2)GlcAOMe | Rha(a1-2)Glc | Rha(a1-2)Glc(b1-2)Glc | Rha(a1-2)Glc(b1-2)GlcA | Rha(a1-2)Glc(b1-2)GlcA6Me | Rha(a1-2)Glc(b1-2)GlcAOMe | Rha(a1-2)Glc(b1-6)Glc | Rha(a1-2)GlcA(b1-2)GlcA | Rha(a1-2)GlcAOMe(b1-2)GlcAOMe | Rha(a1-2)Rha(a1-2)Gal(b1-4)[Glc(b1-2)]GlcA | Rha(a1-2)Xyl | Rha(a1-2)Xyl(b1-2)Glc | Rha(a1-2)Xyl(b1-2)GlcA | Rha(a1-2)Xyl(b1-2)GlcA6Me | Rha(a1-2)Xyl(b1-2)GlcAOMe | Rha(a1-2)Xyl3Ac | Rha(a1-2)Xyl4Ac | Rha(a1-2)[Glc(b1-3)]Glc | Rha(a1-2)[Glc(b1-6)]Gal(b1-2)GlcA6Me | Rha(a1-2)[Rha(a1-4)]Glc | Rha(a1-2)[Rha(a1-6)]Gal | Rha(a1-2)[Rha(a1-6)]Glc | Rha(a1-2)[Xyl(b1-4)]Glc | Rha(a1-2)[Xyl(b1-4)]Glc(b1-6)Glc | Rha(a1-3)GlcA | Rha(a1-3)[Rha(a1-4)]Gal | Rha(a1-4)Gal(b1-2)GlcA | Rha(a1-4)Gal(b1-2)GlcAOMe | Rha(a1-4)Gal(b1-2)GlcOMe | Rha(a1-4)Gal(b1-4)Gal(b1-4)GalGro | Rha(a1-4)Xyl(b1-2)Glc | Rha(a1-4)Xyl(b1-2)GlcA | Rha(a1-4)Xyl(b1-2)GlcAOMe | Rha(a1-6)Glc | Rha(a1-6)[Xyl(b1-3)Xyl(b1-2)]Glc(b1-2)Glc | Rha(b1-2)Glc(b1-2)GlcA | Rha1Fer(a1-4)Fruf(b2-1)GlcOBz | RhaOMe(a1-2)[RhaOMe(a1-6)]GlcOMe-ol | RhaOMe(a1-6)GlcOMe(b1-2)GlcOMe-ol | Xyl(a1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(a1-3)Fuc(a1-4)Rha | Xyl(a1-6)Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc-ol | Xyl(b1-2)Ara(a1-6)Glc | Xyl(b1-2)Ara(a1-6)GlcNAc | Xyl(b1-2)Ara(a1-6)[Glc(b1-2)]Glc | Xyl(b1-2)Ara(a1-6)[Glc(b1-4)]GlcNAc | Xyl(b1-2)D-Fuc(b1-6)Glc | Xyl(b1-2)D-Fuc(b1-6)GlcNAc | Xyl(b1-2)D-Fuc(b1-6)[Glc(b1-2)]Glc | Xyl(b1-2)Fuc(a1-6)Glc | Xyl(b1-2)Fuc(a1-6)GlcNAc | Xyl(b1-2)Fuc(b1-6)Glc | Xyl(b1-2)Fuc(b1-6)GlcNAc | Xyl(b1-2)Fuc(b1-6)[Glc(b1-2)]Glc | Xyl(b1-2)Gal(b1-2)GlcA6Me | Xyl(b1-2)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)Rha(a1-2)Ara | Xyl(b1-2)Xyl(b1-3)[Rha(b1-2)Rha(b1-4)]Xyl | Xyl(b1-2)[Glc(b1-3)]Ara | Xyl(b1-2)[Glc2Ac(b1-4)]Glc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(a1-6)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc-ol | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)ManNAc | Xyl(b1-2)[Man(a1-6)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Rha(a1-3)]GlcA | Xyl(b1-3)Ara | Xyl(b1-3)Xyl(b1-2)[Rha(a1-6)]Glc(b1-2)Glc | Xyl(b1-3)Xyl(b1-4)Rha(a1-2)[Rha(a1-6)]Glc | Xyl(b1-3)Xyl(b1-4)Rha(a1-2)[Rha(a1-6)]Glc(b1-2)Glc | Xyl(b1-4)Rha(a1-2)Ara | Xyl(b1-4)Rha(a1-2)D-Fuc | Xyl(b1-4)Rha(a1-2)D-FucOMe | Xyl(b1-4)Rha(a1-2)Fuc | Xyl(b1-4)Rha(a1-2)Fuc3Ac | Xyl(b1-4)Rha(a1-2)Fuc4Ac | Xyl(b1-4)Rha(a1-2)Glc | Xyl(b1-4)Rha(a1-2)[Rha(a1-3)]Fuc4Ac | Xyl(b1-4)Rha(a1-2)[Rha(a1-6)]Glc | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl3Ac(b1-4)Xyl(b1-4)Xyl(b1-4)[GlcA(a1-2)]Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl3Ac(b1-4)Xyl(b1-4)Xyl(b1-4)[GlcA(a1-2)]Xyl3Ac(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl3Ac(b1-4)Xyl(b1-4)Xyl(b1-4)[GlcA4Me(a1-2)]Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl3Ac(b1-4)Xyl(b1-4)Xyl(b1-4)[GlcA4Me(a1-2)]Xyl3Ac(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)[GlcA(a1-2)]Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)[GlcAOMe(a1-2)]Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl2Ac3Ac4Ac(b1-3)Ara | Xyl4Ac(b1-3)Ara | XylOMe(b1-2)[RhaOMe(a1-6)]GlcOMe(b1-2)GlcOMe-ol | XylOMe(b1-3)XylOMe(b1-2)[RhaOMe(a1-6)]GlcOMe(b1-2)GlcOMe-ol | XylOMe(b1-4)RhaOMe(a1-2)D-FucOMe | XylOMe(b1-4)RhaOMe(a1-2)[RhaOMe(a1-6)]GlcOMe | XylOMe(b1-4)RhaOMe(a1-2)[RhaOMe(a1-6)]GlcOMe-ol | Xylf(b1-2)Xyl(b1-3)[Rha(b1-2)Rha(b1-4)]Xyl | [Araf(a1-3)Gal(b1-3)Gal(b1-6)]Gal(b1-3)Gal | [Araf(a1-3)Gal(b1-6)]Gal(b1-3)Gal | [Gal(a1-4)Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)[Man(b1-4)Man(b1-4)Man(b1-4)Gal(a1-6)]Man(b1-2)[Gal(a1-6)]Man(b1-2)[Gal(a1-4)Gal(a1-6)]Man(b1-4)Man | [Gal(a1-6)]Man(b1-4)Man | [Gal(a1-6)]Man(b1-4)Man(b1-4)Man | [Gal(a1-6)]Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | [Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)Man(b1-4)Man | [Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)[Gal(a1-6)]Man(b1-4)Man | [Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Gal(b1-3)Gal(b1-6)[Araf(a1-3)]Gal(b1-6)]Gal(b1-3)Gal | [Gal(b1-3)Gal(b1-6)]Gal(b1-3)Gal | [Gal(b1-6)Gal(b1-6)Gal(b1-6)]Gal(b1-3)Gal | [Gal(b1-6)Gal(b1-6)]Gal(b1-3)Gal | [Gal(b1-6)]Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Araf(a1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Araf(a1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Araf(a1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Araf(a1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-5)Araf(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-5)Araf(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Fuc(a1-2)Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-5)Araf(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-5)Araf(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Gal(b1-2)Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc | [Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)[Gal(b1-5)Araf(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc(b1-4)Glc |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Family | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fabaceae | 1 | 4 | 1 | 3 | 1 | 1 | 1 | 0 | 1 | 3 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 4 | 2 | 1 | 1 | 2 | 2 | 7 | 7 | 4 | 4 | 4 | 4 | 4 | 2 | 8 | 4 | 2 | 5 | 4 | 2 | 2 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 3 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 2 | 5 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 3 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 3 | 1 | 5 | 0 | 0 | 1 | 3 | 1 | 1 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 4 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | 0 | 1 | 1 | 1 | 1 | 5 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 3 | 2 | 0 | 0 | 0 | 1 | 1 | 4 | 6 | 1 | 1 | 1 | 1 | 3 | 4 | 2 | 1 | 1 | 1 | 4 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 7 | 2 | 5 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 5 | 1 | 1 | 1 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 4 | 1 | 1 | 1 | 1 | 1 | 5 | 1 | 11 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 4 | 6 | 4 | 4 | 4 | 1 | 1 | 5 | 4 | 1 | 4 | 1 | 1 | 0 | 1 | 1 | 1 | 7 | 1 | 1 | 2 | 3 | 23 | 6 | 7 | 0 | 1 | 9 | 3 | 4 | 1 | 3 | 1 | 1 | 1 | 1 | 3 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 0 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 2 | 0 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 2 | 1 | 2 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 7 | 1 | 1 | 1 | 2 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 3 | 16 | 18 | 2 | 1 | 2 | 1 | 9 | 13 | 2 | 1 | 1 | 3 | 2 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 4 | 1 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| Fagaceae | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Polygalaceae | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 2 | 1 | 2 | 2 | 1 | 2 | 1 | 1 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Quillajaceae | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Determines whether glycan belongs to a specified class
Convert list of IUPAC-condensed glycans to isomeric SMILES using GlyLES
['O1C(O)[C@H](O)[C@@H](O)[C@H](O[C@@H]2O[C@H](CO)[C@H](O)[C@H](O[C@]3(C(=O)O)C[C@H](O)[C@@H](NC(C)=O)[C@H]([C@H](O)[C@H](O)CO)O3)[C@H]2O)[C@H]1CO']Converts composition from any common format to standardized dictionary
{'HexNAc': 2, 'Hex': 1, 'dHex': 3, 'Neu5Ac': 1}
{'HexNAc': 2, 'Hex': 1, 'dHex': 3, 'Neu5Ac': 1}Convert glycan from IUPAC-extended, LinearCode, GlycoCT, WURCS, Oxford, GLYCAM, GlycoWorkBench, CSDB-linear, KCF, GlyConnect IDs, and GlyTouCanIDs to standardized IUPAC-condensed format
print(canonicalize_iupac("NeuAc?1-36SGalb1-4GlcNACb1-6(Fuc?1-2Galb1-4GlcNacb1-3Galb1-3)GalNAc-sp3"))
print(canonicalize_iupac("WURCS=2.0/5,11,10/[a2122h-1b_1-5_2*NCC/3=O][a1122h-1b_1-5][a1122h-1a_1-5][a2112h-1b_1-5][a1221m-1a_1-5]/1-1-2-3-1-4-3-1-4-5-5/a4-b1_a6-k1_b4-c1_c3-d1_c6-g1_d2-e1_e4-f1_g2-h1_h4-i1_i2-j1"))
print(canonicalize_iupac("Ma3(Ma6)Mb4GNb4GN;N"))
print(canonicalize_iupac("α-D-Manp-(1→3)[α-D-Manp-(1→6)]-β-D-Manp-(1→4)-β-D-GlcpNAc-(1→4)-β-D-GlcpNAc-(1→"))
print(canonicalize_iupac("""RES
1b:b-dgal-HEX-1:5
2s:n-acetyl
3b:b-dgal-HEX-1:5
4b:b-dglc-HEX-1:5
5b:b-dgal-HEX-1:5
6b:a-dglc-HEX-1:5
7b:b-dgal-HEX-1:5
8b:a-lgal-HEX-1:5|6:d
9b:a-dgal-HEX-1:5
10s:n-acetyl
11s:n-acetyl
12b:b-dglc-HEX-1:5
13b:b-dgal-HEX-1:5
14b:a-lgal-HEX-1:5|6:d
15b:a-lgal-HEX-1:5|6:d
16s:n-acetyl
17s:n-acetyl
18b:b-dgal-HEX-1:5
LIN
1:1d(2+1)2n
2:1o(3+1)3d
3:3o(3+1)4d
4:4o(-1+1)5d
5:5o(-1+1)6d
6:6o(-1+1)7d
7:7o(2+1)8d
8:7o(3+1)9d
9:9d(2+1)10n
10:6d(2+1)11n
11:5o(-1+1)12d
12:12o(-1+1)13d
13:13o(2+1)14d
14:12o(-1+1)15d
15:12d(2+1)16n
16:4d(2+1)17n
17:1o(6+1)18d
"""))Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc(b1-6)]GalNAc
Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc
Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc
Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc
Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-?)GlcNAc(a1-?)[Fuc(a1-2)Gal(b1-?)[Fuc(a1-?)]GlcNAc(b1-?)]Gal(b1-?)GlcNAc(b1-3)Gal(b1-3)[Gal(b1-6)]GalNAc
def get_possible_linkages(
wildcard:str, # Pattern to match, ? can be wildcard
linkage_list:list={'b2-4', 'b1-6', '?1-?', 'b2-7', 'a2-3', 'a1-1', 'b1-5', 'a2-11', 'b2-8', 'b1-4', 'a1-3', 'b2-2', 'b1-2', 'b1-3', 'a2-1', 'a1-5', '?2-6', 'a1-8', 'a1-11', '?1-3', '?1-6', 'a2-4', 'a1-2', 'a2-2', 'b2-6', 'b2-1', 'a2-8', 'b1-?', '?1-2', '?2-3', '?2-8', 'a2-6', 'a1-7', 'b1-7', '?2-?', 'a1-6', 'a2-9', 'a1-9', 'a2-?', 'b2-5', 'a1-4', 'b1-9', '1-6', 'a2-7', 'b1-1', 'a1-?', 'b1-8', 'b2-3', 'a2-5', '?1-4', '1-4'}, # List of linkages to search
)->set: # Matching linkages
Retrieves all linkages that match a given wildcard pattern
{'a1-1',
'a1-2',
'a1-3',
'a1-4',
'a1-5',
'a1-6',
'a1-7',
'a1-8',
'a1-9',
'a1-?'}Retrieves all matching common monosaccharides of a type
Check whether two repeat units could stem from the same repeating structure
Determines glycan class
for interacting with the databases contained in glycowork, delivering insights for sequences of interest
Print meta-information about a glycan
Test get_insight with 'Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc'
Let's get rolling! Give us a few moments to crunch some numbers.
This glycan occurs in the following species: ['Acanthocheilonema_viteae', 'Adeno-associated_dependoparvovirusA', 'Aedes_aegypti', 'Angiostrongylus_cantonensis', 'Anopheles_gambiae', 'Antheraea_pernyi', 'Apis_mellifera', 'Ascaris_suum', 'Autographa_californica_nucleopolyhedrovirus', 'AvianInfluenzaA_Virus', 'Bombus_ignitus', 'Bombyx_mori', 'Bos_taurus', 'Brugia_malayi', 'Caenorhabditis_elegans', 'Cardicola_forsteri', 'Cooperia_onchophora', 'Cornu_aspersum', 'Crassostrea_gigas', 'Crassostrea_virginica', 'Cricetulus_griseus', 'Danio_rerio', 'Dictyocaulus_viviparus', 'Dirofilaria_immitis', 'Drosophila_melanogaster', 'Fasciola_hepatica', 'Gallus_gallus', 'Glossina_morsitans', 'Haemonchus_contortus', 'Haliotis_tuberculata', 'Heligmosomoides_polygyrus', 'Helix_lucorum', 'Homo_sapiens', 'HumanImmunoDeficiency_Virus', 'Hylesia_metabus', 'Hypsibius_exemplaris', 'Lutzomyia_longipalpis', 'Lymantria_dispar', 'Macaca_mulatta', 'Mamestra_brassicae', 'Megathura_crenulata', 'Mus_musculus', 'Nilaparvata_lugens', 'Oesophagostomum_dentatum', 'Onchocerca_volvulus', 'Onchocerca_volvulus', 'Ophiactis_savignyi', 'Opisthorchis_viverrini', 'Ostrea_edulis', 'Ovis_aries', 'Pan_troglodytes', 'Pristionchus_pacificus', 'Ramazzottius_varieornatus', 'Rattus_norvegicus', 'Schistosoma_mansoni', 'SemlikiForest_Virus', 'Spodoptera_frugiperda', 'Sus_scrofa', 'Tick_borne_encephalitis_virus', 'Tribolium_castaneum', 'Trichinella_spiralis', 'Trichoplusia_ni', 'Trichuris_suis', 'Tropidolaemus_subannulatus', 'Volvarina_rubella', 'undetermined', 'unidentified_influenza_virus']
Puh, that's quite a lot! Here are the phyla of those species: ['Arthropoda', 'Artverviricota', 'Chordata', 'Cossaviricota', 'Echinodermata', 'Kitrinoviricota', 'Mollusca', 'Negarnaviricota', 'Nematoda', 'Platyhelminthes', 'Tardigrada', 'Virus']
This glycan contains the following motifs: ['Chitobiose', 'Trimannosylcore', 'core_fucose']
This is the GlyTouCan ID for this glycan: G63041RA
This glycan has been reported to be expressed in: ['2A3_cell_line', 'A549_cell_line', 'AML_193_cell_line', 'C10_cell_line', 'CHOK1_cell_line', 'CHOS_cell_line', 'COLO_205_cell_line', 'COLO_320_cell_line', 'CRL_1620_cell_line', 'Caco_2_cell_line', 'Cal-27_cell_line', 'Cervicovaginal_Secretion', 'Co_115_cell_line', 'EOL_1_cell_line', 'FaDu_cell_line', 'HCT_15_cell_line', 'HCT_8_cell_line', 'HEK293_cell_line', 'HEL92_1_7_cell_line', 'HEL_cell_line', 'HL_60_cell_line', 'HT_29_cell_line', 'KG_1_cell_line', 'KG_1a_cell_line', 'KM12_cell_line', 'Kasumi_1_cell_line', 'LS174T_cell_line', 'LS180_cell_line', 'LS411N_cell_line', 'LoVo_cell_line', 'MDA_MB_231BR_cell_line', 'ME_1_cell_line', 'ML_1_cell_line', 'MOLM_13_cell_line', 'MOLM_14_cell_line', 'MV4_11_cell_line', 'M_07e_cell_line', 'NB_4_cell_line', 'NS0_cell_line', 'OCI_AML2_cell_line', 'OCI_AML3_cell_line', 'PLB_985_cell_line', 'RKO_cell_line', 'SCC-9_cell_line', 'SCC_25_cell_line', 'SW1116_cell_line', 'SW1398_cell_line', 'SW1463_cell_line', 'SW480_cell_line', 'SW48_cell_line', 'SW620_cell_line', 'SW948_cell_line', 'T84_cell_line', 'TF_1_cell_line', 'THP_1_cell_line', 'U_937_cell_line', 'VU-147T_cell_line', 'WiDr_cell_line', 'alveolus_of_lung', 'brain', 'brain', 'cerebellar_cortex', 'cerebellar_cortex', 'cerebellar_cortex', 'cerebellar_cortex', 'cerebellum', 'colon', 'cortex', 'digestive_tract', 'digestive_tract', 'forebrain', 'gills', 'gills', 'heart', 'heart', 'heart', 'hindbrain', 'hippocampal_formation', 'hippocampus', 'hippocampus', 'hippocampus', 'hippocampus', 'iPS1A_cell_line', 'iPS2A_cell_line', 'kidney', 'liver', 'liver', 'liver', 'lung', 'mantle', 'mantle', 'metastatic_pancreatic_ductal_adenocarcinoma', 'milk', 'mucus', 'muscle_of_leg', 'nerve_ending', 'ovary', 'pancreas', 'placenta', 'prefrontal_cortex', 'prefrontal_cortex', 'prefrontal_cortex', 'prefrontal_cortex', 'primary_pancreatic_ductal_adenocarcinoma', 'prostate_gland', 'seminal_fluid', 'striatum', 'striatum', 'striatum', 'striatum', 'testicle', 'testis', 'trachea', 'urine', 'urothelium']
This glycan has been reported to be dysregulated in (disease, direction, sample): [('REM_sleep_behavior_disorder', 'down', 'serum'), ('benign_breast_tumor_tissues_vs_para_carcinoma_tissues', 'up', 'breast'), ('cystic_fibrosis', 'up', 'sputum'), ('female_breast_cancer', 'up', 'breast'), ('female_breast_cancer', 'up', 'cell_line'), ('prostate_cancer', 'up', 'prostate_cancer_biopsy'), ('thyroid_gland_papillary_carcinoma', 'up', 'serum'), ('urinary_bladder_cancer', 'down', 'urine')]
That's all we can do for you at this point!Convert between GlyTouCan IDs and IUPAC-condensed glycans
for performing regular expression-like searches in glycans, very powerful to find complicated motifs
Find matches for glyco-regular expression in glycan
# {} = between min and max occurrences, e.g., "Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc"
# * = zero or more occurrences, e.g., "Hex-HexNAc-([Hex|Fuc])*-HexNAc"
# + = one or more occurrences, e.g., "Hex-HexNAc-([Hex|Fuc])+-HexNAc"
# ? = zero or one occurrence, e.g., "Hex-HexNAc-([Hex|Fuc])?-HexNAc"
# {1,} = at minimum one occurrence, e.g., "Hex-HexNAc-([Hex|Fuc]){1,}-HexNAc"
# {,1} = at maximum one occurrence, e.g., "Hex-HexNAc-([Hex|Fuc]){,1}-HexNAc"
# {2} = exactly two occurrences, e.g., "Hex-HexNAc-([Hex|Fuc]){2}-HexNAc"
# ^ = start of sequence, e.g., "^Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc"
# % = middle of sequence (i.e., neither start nor end)
# $ = end of sequence, e.g., "Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc$"
# ?<= = lookbehind (i.e., provided pattern must be present before rest of pattern but is not included in match), e.g., "(?<=Xyl-)Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc"
# ?<! = negative lookbehind (i.e., provided pattern is not present before rest of pattern and is also not included in match), e.g., "(?<!Xyl-)Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc"
# ?= = lookahead (i.e., provided pattern must be present after rest of pattern but is not included in match), e.g., "Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc(?=-HexNAc)"
# ?! = negative lookahead (i.e., provided pattern is not present after rest of pattern and is not included in match), e.g., "Hex-HexNAc-([Hex|Fuc]){1,2}-HexNAc(?!-HexNAc)"
# Example: extracting the sequence from the a1-6 branch of N-glycans
pattern = "r[Sia]{,1}-Monosaccharide-([dHex]){,1}-Monosaccharide(?=-Mana6-Monosaccharide)"
print(get_match(pattern, "GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc"))
print(get_match(pattern, "GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc"))
print(get_match(pattern, "GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc"))
print(get_match(pattern, "GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)GalNAc(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc"))['Gal(b1-4)GlcNAc']
['GalNAc(b1-4)GlcNAc']
['Neu5Ac(a2-6)GalNAc(b1-4)GlcNAc']
['Neu5Gc(a2-6)GalNAc(b1-4)[Fuc(a1-3)]GlcNAc']For interested users, we here compile a selection of regular expression patterns that we find useful in our own work:
Find glyco-regular expression matches in list of glycans
Convert glycan motif to regular expression pattern
helper functions to map m/z–>composition, composition–>structure, structure–>motif, and more
Tokenize word by indexing characters in library
Pad sequences to same length using padding token
[None, None, None, None, None, 25, 25]Remove modifications from all monosaccharides in glycan
'Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc'
def stemify_dataset(
df:DataFrame, # DataFrame with glycan column
stem_lib:dict[str, str] | None=None, # Modified to core monosaccharide mapping; default:created from lib
libr:dict[str, int] | None=None, # Glycoletter to index mapping
glycan_col_name:str='glycan', # Column name for glycans
rarity_filter:int=1, # Minimum occurrences to keep modification
)->DataFrame: # DataFrame with stemified glycans
Remove monosaccharide modifications from all glycans in dataset
def mask_rare_glycoletters(
glycans:list, # List of IUPAC-condensed glycans
thresh_monosaccharides:int | None=None, # Threshold for rare monosaccharides (default: 0.001*len(glycans))
thresh_linkages:int | None=None, # Threshold for rare linkages (default: 0.03*len(glycans))
)->list: # List of glycans with masked rare elements
Mask rare monosaccharides and linkages in glycans
def mz_to_composition(
mz_value:float, # m/z value from mass spec
mode:str='negative', # MS mode: positive/negative
mass_value:str='monoisotopic', # Mass type: monoisotopic/average
modification:str | None=None, # Reducing end modification: reduced/2AA/2AB
sample_prep:str='underivatized', # Sample preparation method: underivatized/permethylated/peracetylated
mass_tolerance:float=0.5, # Mass tolerance for matching
kingdom:str='Animalia', # Taxonomic kingdom filter for choosing a subset of glycans to consider
glycan_class:str='all', # Glycan class: N/O/lipid/free/all
df_use:pandas.DataFrame | None=None, # Custom glycan database
filter_out:set[str] | None=None, # Monosaccharides to ignore during composition finding
deprioritized:set[str] | None={'PCho', 'HexA', 'Me'}, # Monosaccharides to use only as fallback if no other composition matches
extras:list=['doubly_charged'], # Additional operations: adduct/doubly_charged
adduct:str | None=None, # Chemical formula of adduct that contributes to m/z, e.g., "C2H4O2"
mass_tag:float | None=None, # Mass in Da of a reducing-end label (e.g., 137.14 for 2AA, 219.21 for 2AB+procA), subtracted from mz_value
)->list: # List of matching compositions
Map m/z value to matching monosaccharide composition
[{'Hex': 2, 'HexNAc': 2, 'Neu5Ac': 2}]
def match_composition_relaxed(
composition:dict, # Dictionary indicating composition (e.g. {"dHex": 1, "Hex": 1, "HexNAc": 1})
glycan_class:str='N', # Glycan class: N/O/lipid/free
kingdom:str='Animalia', # Taxonomic kingdom filter for choosing a subset of glycans to consider
df_use:pandas.DataFrame | None=None, # Custom glycan database
)->list: # List of matching glycans
Map coarse-grained composition to matching glycans
['Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc',
'Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc',
'Gal(b1-4)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)]GalNAc',
'Gal(?1-3/4)Gal(b1-3/4)[Fuc(a1-3/4)]GlcNAc(b1-6)[Gal(b1-3)]GalNAc',
'Gal(b1-4)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-3)GalNAc',
'Fuc(a1-2)Gal(b1-3/4)GlcNAc(b1-3)Gal(b1-3)[Gal(b1-6)]GalNAc',
'Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[Gal(?1-?)Gal(b1-3)]GalNAc',
'Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-3)GalNAc',
'Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(?1-3/4)Gal(b1-3)GalNAc',
'Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal',
'Gal(?1-?)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)]GalNAc',
'Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[Gal(a1-3)Gal(b1-3)]GalNAc',
'Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)]GalNAc',
'Fuc(a1-2)Gal(b1-3)Gal(b1-3)GlcNAc(b1-6)[Gal(b1-3)]GalNAc',
'Fuc(a1-2)Gal(b1-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc',
'Gal(b1-4)Gal(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc',
'Fuc(a1-2)Gal(?1-?)Gal(b1-3/4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc',
'Gal(a1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-3)Gal(b1-3)GalNAc',
'Man(a1-6)Glc(a1-4)GlcNAc(b1-4)[Fuc(a1-2)]Gal(b1-3)GalNAc',
'Man(a1-6)Glc(b1-4)GlcNAc(b1-4)[Fuc(a1-2)]Gal(b1-3)GalNAc',
'Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-?)Man',
'Gal(b1-2)Gal(a1-3)[Fuc(a1-2)]Gal(b1-3)[GlcNAc(b1-6)]GalNAc',
'Fuc(a1-2)Gal(a1-3)Gal(a1-4)Gal(b1-3)[GlcNAc(b1-6)]GalNAc',
'Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)]Gal(b1-3)GalNAc',
'Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-?)Gal(b1-6)[Gal(b1-3)]GalNAc']Find minimum set of glycans characterizing matched composition
['Neu5Ac(a2-3)Gal(b1-3)GalNAc', 'Gal(b1-3)[Neu5Ac(a2-6)]GalNAc', '{Neu5Ac(a2-3/6)}Gal(b1-3)GalNAc', 'Neu5Ac(a2-3)[GalNAc(b1-4)]Gal', 'Gal(a1-3)[Neu5Ac(a2-6)]GalNAc', 'Neu5Ac(a2-3/6)Gal(b1-3)GalNAc', 'Neu5Ac(a2-6)Gal(b1-3)GalNAc', 'Gal(?1-3)[Neu5Ac(a2-6)]GalNAc', 'Neu5Ac(a2-3/6)Gal(?1-3)GalNAc', 'Neu5Ac(a2-?)Hex(?1-?)GalNAc', 'Neu5Ac(a2-3)Gal(?1-?)GalNAc', 'Neu5Ac(a2-3/6)GalNAc(a1-6)Gal', 'Neu5Ac(a2-6)Gal(a1-3)GalNAc', 'Gal(b1-4)[Neu5Ac(a2-6)]GalNAc', 'Neu5Ac(a2-3)GalNAc(b1-3)Gal']['Neu5Ac(a2-3)Gal(b1-3)GalNAc',
'Neu5Ac(a2-3/6)Gal(b1-3)GalNAc',
'Gal(b1-3)[Neu5Ac(a2-6)]GalNAc',
'Gal(a1-3)[Neu5Ac(a2-6)]GalNAc',
'{Neu5Ac(a2-3/6)}Gal(b1-3)GalNAc',
'Neu5Ac(a2-3)[GalNAc(b1-4)]Gal',
'Neu5Ac(a2-6)Gal(b1-3)GalNAc',
'Neu5Ac(a2-3/6)GalNAc(a1-6)Gal',
'Neu5Ac(a2-6)Gal(a1-3)GalNAc',
'Gal(b1-4)[Neu5Ac(a2-6)]GalNAc',
'Neu5Ac(a2-3)GalNAc(b1-3)Gal']
def mz_to_structures(
mz_list:list, # List of precursor masses
glycan_class:str, # Glycan class: N/O/lipid/free
kingdom:str='Animalia', # Taxonomic kingdom filter for choosing a subset of glycans to consider
abundances:pandas.DataFrame | None=None, # Sample abundances matrix
mode:str='negative', # MS mode: positive/negative
mass_value:str='monoisotopic', # Mass type: monoisotopic/average
sample_prep:str='underivatized', # Sample prep: underivatized/permethylated/peracetylated
mass_tolerance:float=0.5, # Mass tolerance for matching
modification:str | None=None, # Reducing end modification: reduced/2AA/2AB
df_use:pandas.DataFrame | None=None, # Custom glycan database
filter_out:set[str] | None=None, # Monosaccharides to ignore
deprioritized:set[str] | None={'PCho', 'HexA', 'Me'}, # Monosaccharides to use only as fallback if no other composition matches
verbose:bool=False, # Whether to print non-matching compositions
mass_tag:float | None=None, # Mass in Da of a reducing-end label (e.g., 137.14 for 2AA), subtracted from each m/z before matching
)->pandas.DataFrame | list: # DataFrame of structures x intensities or empty list
Map precursor masses to structures, supporting accompanying relative intensities
0 compositions could not be matched. Run with verbose = True to see which compositions.| glycan | abundance | |
|---|---|---|
| 0 | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc | 0 |
def compositions_to_structures(
composition_list:list, # List of compositions like {'Hex': 1, 'HexNAc': 1}
glycan_class:str='N', # Glycan class: N/O/lipid/free
kingdom:str='Animalia', # Taxonomic kingdom filter for choosing a subset of glycans to consider
abundances:pandas.DataFrame | None=None, # Sample abundances matrix
df_use:pandas.DataFrame | None=None, # Custom glycan database
verbose:bool=False, # Whether to print non-matching compositions
)->DataFrame: # DataFrame of structures x intensities
Map compositions to structures, supporting accompanying relative intensities
0 compositions could not be matched. Run with verbose = True to see which compositions.| glycan | abundance | |
|---|---|---|
| 0 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 0 |
| 1 | Neu5Ac(a2-8)Neu5Ac(a2-6)[Gal(b1-3)]GalNAc | 0 |
| 2 | Neu5Ac(a2-3)[Neu5Ac(a2-6)]Gal(b1-3)GalNAc | 0 |
| 3 | Neu5Ac(a2-3)Gal(b1-4)[Neu5Ac(a2-6)]GalNAc | 0 |
0 compositions could not be matched. Run with verbose = True to see which compositions.| glycan | abundance | |
|---|---|---|
| 0 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 0 |
| 1 | Neu5Ac(a2-8)Neu5Ac(a2-6)[Gal(b1-3)]GalNAc | 0 |
| 2 | Neu5Ac(a2-3)[Neu5Ac(a2-6)]Gal(b1-3)GalNAc | 0 |
| 3 | Neu5Ac(a2-3)Gal(b1-4)[Neu5Ac(a2-6)]GalNAc | 0 |
Convert glycan structure to base topology
'Neu5Ac(?1-?)HexOS(?1-?)[Neu5Ac(?1-?)]HexNAc'Map glycan to its composition
{'Hex': 1, 'HexNAc': 1, 'Neu5Ac': 2, 'S': 1}
def glycan_to_mass(
glycan:str, # Glycan in IUPAC-condensed format
mass_value:str='monoisotopic', # Mass type: monoisotopic/average
sample_prep:str='underivatized', # Sample prep: underivatized/permethylated/peracetylated
stem_libr:dict[str, str] | None=None, # Modified to core monosaccharide mapping
adduct:str | float | None=None, # Chemical formula of adduct (e.g., "C2H4O2") OR its exact mass in Da
modification:str | None=None, # Reducing end modification: reduced/2AA/2AB
)->float: # Theoretical mass
Calculate theoretical mass from glycan
def composition_to_mass(
dict_comp_in:dict, # Composition dictionary of monosaccharide:count
mass_value:str='monoisotopic', # Mass type: monoisotopic/average
sample_prep:str='underivatized', # Sample prep: underivatized/permethylated/peracetylated
adduct:str | float | None=None, # Chemical formula of adduct (e.g., "C2H4O2") OR its exact mass in Da
modification:str | None=None, # Reducing end modification: reduced/2AA/2AB
)->float: # Theoretical mass
Calculate theoretical mass from composition
Calculate mass of adduct from chemical formula, including signed formulas
def get_unique_topologies(
composition:dict, # Composition dictionary of monosaccharide:count
glycan_type:str, # Glycan class: N/O/lipid/free/repeat
df_use:pandas.DataFrame | None=None, # Custom glycan database to use for mapping
universal_replacers:dict[str, str] | None=None, # Base-to-specific monosaccharide mapping
taxonomy_rank:str='Kingdom', # Taxonomic rank for filtering
taxonomy_value:str='Animalia', # Value at taxonomy rank
)->list: # List of unique base topologies
Get all observed unique base topologies for composition
['Hex(?1-?)HexNAc(?1-?)HexNAc',
'HexNAc(?1-?)HexNAc(?1-?)Hex',
'HexNAc(?1-?)[HexNAc(?1-?)]Hex',
'Hex(?1-?)[HexNAc(?1-?)]HexNAc',
'HexNAc(?1-?)Hex(?1-?)HexNAc']
def get_random_glycan(
n:int=1, # How many random glycans to sample
glycan_class:str='all', # Glycan class: N/O/lipid/free/repeat/all
kingdom:str='Animalia', # Taxonomic kingdom filter for choosing a subset of glycans to consider
)->str | list[str]: # Returns a random glycan or list of glycans if n > 1
Sample random glycans from the SugarBase database