df_fish = df_species[df_species.Class == 'Actinopterygii'].reset_index(drop = True)Examples
On this page, there are different toy examples demonstrating how you can apply glycowork to your projects! Here is a summary of the different available case studies:
- Investigation of N-linked and O-linked glycosylation in fish
- Glycan binding specificities of lectins
- Glycan classification using machine learning models
- Constructing and exploring biosynthetic networks
- Deep Learning Code Snippets
- GlycoDraw Code Snippets
Example 1: Investigating N- and O-linked glycosylation in fish
Suppose that after investigating glycans from plants as presented in the core module, you are now wondering how glycans from the Actinopterygii class (ray-finned fish) look like. To satisfy your curiosity, let’s start with importing a dataset from glycowork.glycan_data.loader.df_species and putting a filter for fish glycans.
| Species | Genus | Family | Order | Class | Phylum | Kingdom | Domain | ref | |
|---|---|---|---|---|---|---|---|---|---|
| glycan | |||||||||
| Gal(?1-3/4)[Gal(?1-?)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Acipenser_brevirostrum | Acipenser | Acipenseridae | Acipenseriformes | Actinopterygii | Chordata | Animalia | Eukarya | https://www.frontiersin.org/articles/10.3389/fmolb.2021.778383/full |
| Gal(?1-3/4)[Gal(?1-?)]Gal(b1-4)GlcNAc(b1-2)[Gal(?1-3/4)[Gal(?1-?)]Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Acipenser_brevirostrum | Acipenser | Acipenseridae | Acipenseriformes | Actinopterygii | Chordata | Animalia | Eukarya | https://www.frontiersin.org/articles/10.3389/fmolb.2021.778383/full |
| Gal(?1-3/4)[Gal(?1-?)]Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Acipenser_oxyrinchus | Acipenser | Acipenseridae | Acipenseriformes | Actinopterygii | Chordata | Animalia | Eukarya | https://www.frontiersin.org/articles/10.3389/fmolb.2021.778383/full |
| Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Acipenser_oxyrinchus | Acipenser | Acipenseridae | Acipenseriformes | Actinopterygii | Chordata | Animalia | Eukarya | https://www.frontiersin.org/articles/10.3389/fmolb.2021.778383/full |
| Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3/6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Acipenser_oxyrinchus | Acipenser | Acipenseridae | Acipenseriformes | Actinopterygii | Chordata | Animalia | Eukarya | https://www.frontiersin.org/articles/10.3389/fmolb.2021.778383/full |
To have a better overview of the glycans similarities in fish, you can plot these data using glycowork.motif.analysis.plot_embeddings. The best option here, to visualize the data, is to color the dots by taxonomic family.
plot_embeddings(df_fish.glycan.values.tolist(), label_list = df_fish.Family.values.tolist())Download completed.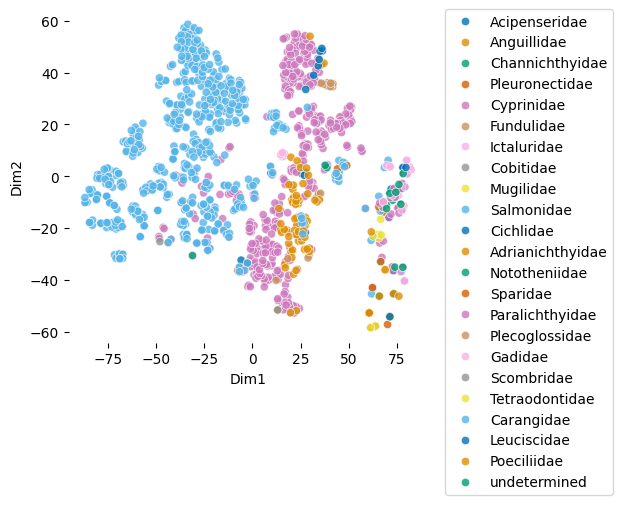
Interesting! First, the graph shows a large Salmonidae cluster in addition to a separate cluster of various families.
Now, let’s focus on further analyses. We will first convert our data into a count table using glycowork.motif.processing.presence_to_matrix and then visualize the results. A heatmap can be generated using glycowork.motif.analysis.get_heatmap, and the yticklabels = True option allows us to display all the labels.
df_map_fish = presence_to_matrix(df_fish, label_col_name = 'Family')
get_heatmap(df_map_fish, motifs = True, feature_set = ['known', 'exhaustive'], datatype = 'presence', show_all = True)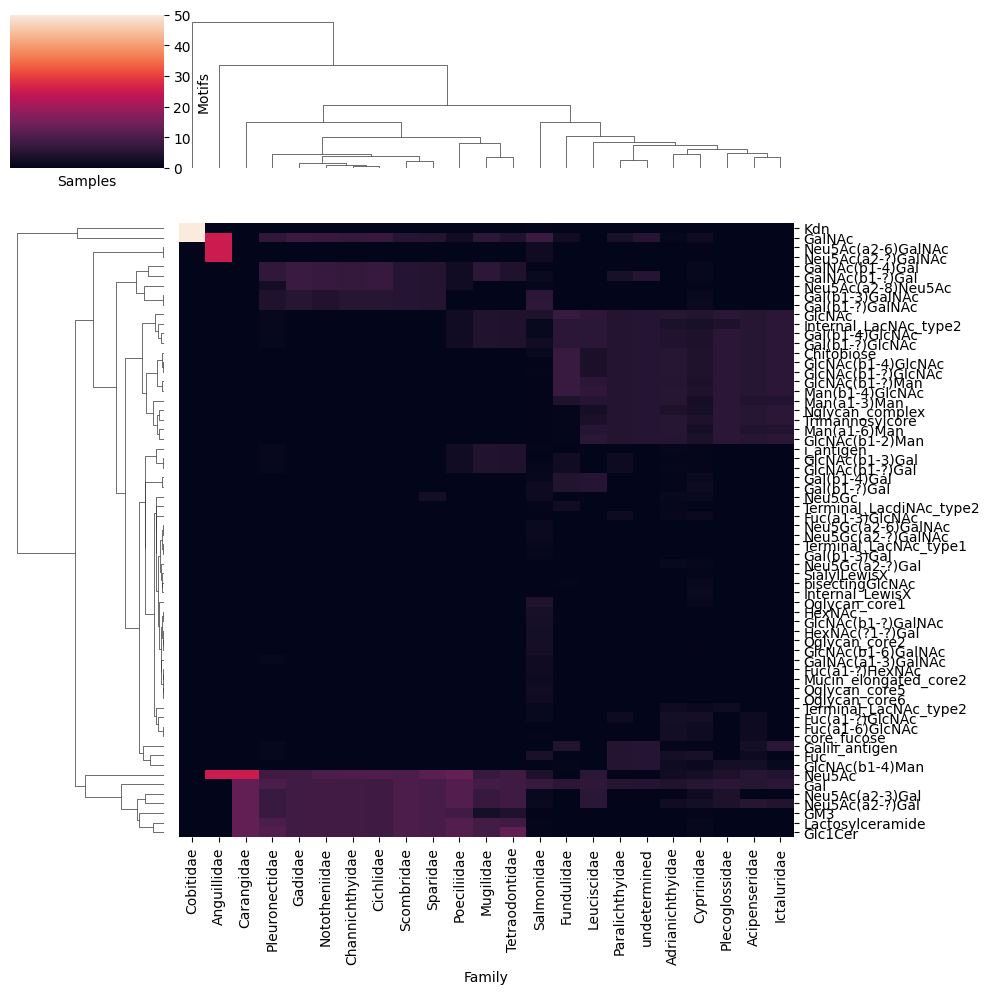
The first thing that catches the eye is the presence of a bunch of glycan motifs in Mugilidae to Cichlidae that are absent in the other classes. One hypothesis for this is that these families mainly contain glycosphingolipids in our database, while the other families focus on N-linked glycans (and Salmonidae + Cyprinidae have both as well-investigated families).
To visualize which dots from the embedding plot correspond to glycosphingolipids, we can focus on 1Cer-containing glycans. We can map them again but with the additional option shape_feature. This parameter allows you to transform dots into crosses if the corresponding glycan contains the specified monosaccharide, modification, or linkage.
plot_embeddings(df_fish.glycan.values.tolist(), label_list = df_fish.Family.values.tolist(), shape_feature = "1Cer")Download completed.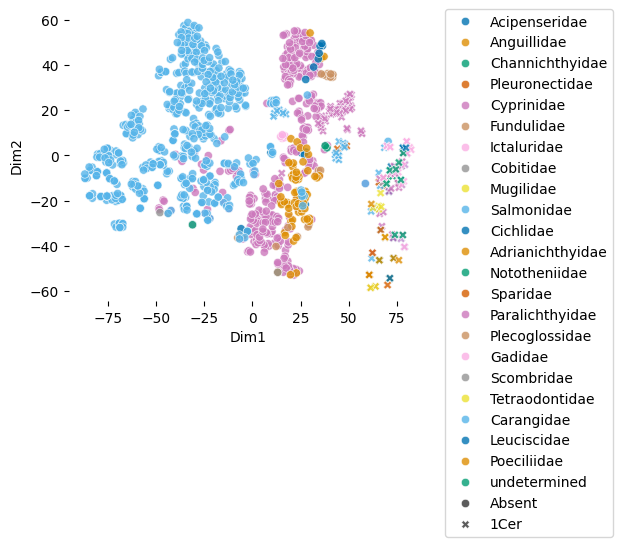
As you can see, in df_species, 1Cer-containing glycans (crosses) are only present in one of the two clusters (in the class Actinopterygii). In this cluster, they in turn show enriched locations, presumably corresponding to certain groups of related glycans.
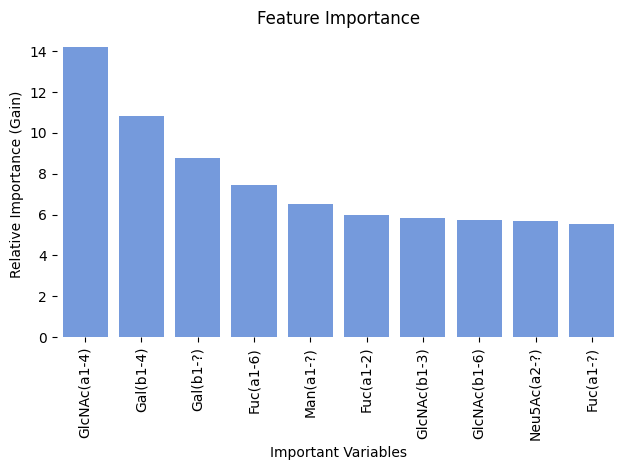
We can also use glycowork to take a look at the overall monosaccharide distribution in fish glycans. glycowork.motif.analysis.characterize_monosaccharide will be our best ally to learn what other monosaccharides, for instance, Gal variants are often connected to.
characterize_monosaccharide('Gal', rank = 'Class', focus = 'Actinopterygii', modifications = True)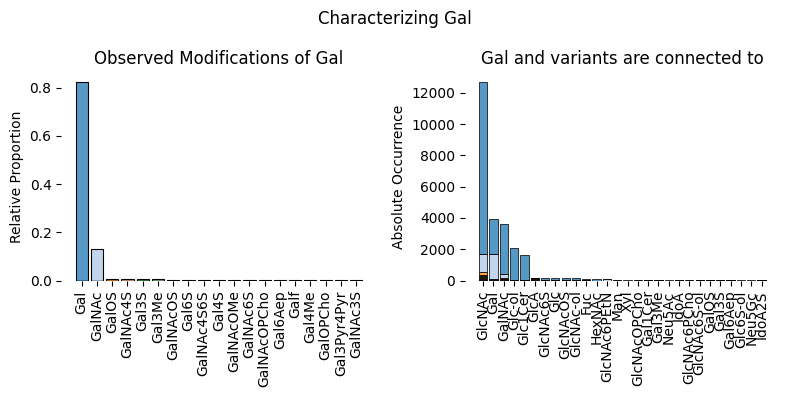
Running this analysis, it appears that Gal is found connected to GlcNAc as well as various other monosaccharides and that it does display several variants, including sulfurylated galactose.
Example 2: Glycan binding specificities of lectins
For this second example, we will focus on a specific set of proteins, called lectins, that bind glycans in a sequence-specific manner. One example for lectins can be found in the hemagglutinin protein of influenza virus. Influenza viruses are well known as the biological agents responsible for the seasonal flu in humans. In general, viruses penetrate inside host cells thanks to a mechanism involving contacts between proteins and glycans. Surface proteins surrounding viruses and eukaryotic cells interact together and allow the entry of viral particles inside the targeted cells. However, different influenza strains may recognize different glycosylations on proteins more or less efficiently.
To measure these glycan binding specificities, one can add virus particles to glycan arrays presenting immobilized glycans. Such a protocol allows screening for specific binding. From such data, we can then ask what are the glycans that are the most efficiently recognized by a given influenza virus?
Fortunately, glycowork can be really helpful to answer such a question! The binding specificity of 1,392 lectins for a vast range of glycans is available in the glycan_binding dataset. Let’s start by importing these data.
| 3-Anhydro-Gal(a1-3)Gal(b1-4)3-Anhydro-Gal(a1-3)Gal4S | 3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S | 3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S | 3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S | 3-Anhydro-Gal(a1-3)Gal4S(b1-4)3-Anhydro-Gal2S(a1-3)Gal4S(b1-4)3-Anhydro-Gal(a1-3)Gal4S | 3dGal | 3dGal(b1-3)[Fuc(a1-4)]Glc | 3dGal(b1-4)Glc | 4d8dNeu5Ac(a2-3)Gal(b1-4)Glc | 4dNeu5Ac(a2-3)Gal(b1-4)Glc | 7dNeu5Ac(a2-3)Gal(b1-4)Glc | 8dNeu5Ac(a2-3)Gal(b1-4)Glc | 8eLeg5Ac7Ac(a2-6)Gal(a1-3)FucNAc(a1-3)GlcNAc(a1-4)8eLeg5Ac7Ac | 8eLeg5Ac7Ac(a2-6)Gal(b1-3)FucNAc(a1-3)GlcNAc(a1-4)8eLeg5Ac7Ac | 9dNeu5Ac(a2-3)Gal(b1-4)Glc | Ara | Ara(a1-2)Xyl(b1-4)Xyl(b1-4)Xyl | Ara(a1-2)[Ara(a1-3)]Xyl(b1-4)Xyl(b1-4)Xyl | Ara(a1-2)[Ara(a1-3)][Xyl(b1-4)]Xyl(b1-4)Xyl(b1-4)Xyl | Ara(a1-3)Xyl(b1-4)Xyl | Ara(a1-3)Xyl(b1-4)Xyl(b1-4)Xyl | Ara(a1-3)[Ara(a1-5)]Ara(a1-5)Ara(a1-5)Ara | Ara(a1-3)[Xyl(b1-4)]Xyl(b1-4)Xyl(b1-4)Xyl | Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara(a1-5)Ara | Ara(a1-5)Ara(a1-5)[Ara(a1-3)]Ara(a1-5)Ara(a1-5)Ara | Araf(a1-3)[Araf(a1-5)]Araf | Araf(a1-3)[Araf(a1-5)]Araf(a1-5)Araf | Araf(a1-5)Araf | Araf(a1-5)Araf(a1-3)[Araf(a1-5)Araf(a1-5)]Araf(a1-5)Araf | Araf(a1-5)Araf(a1-5)Araf(a1-5)Araf(a1-5)Araf(a1-5)Araf | Araf(b1-2)Araf(a1-3)[Araf(b1-2)Araf(a1-5)]Araf(a1-5)Araf | Col(a1-3)[Col(a1-6)]Glc(a1-4)Gal(a1-3)GlcNAc | D-FucNAc | D-Rha | D-Rha4NAc(a1-3)Fuc(a1-4)Glc(b1-3)GalNAc(a1-2)D-Rha4NAc | Fru6P | Fuc | Fuc(a1-2)Gal | Fuc(a1-2)Gal(b1-3)GalNAc | Fuc(a1-2)Gal(b1-3)GalNAc(a1-3)GlcNAc(a1-2)Fuc | Fuc(a1-2)Gal(b1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)GalNAc(b1-3)Gal | Fuc(a1-2)Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)GalNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)GalNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)]Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)GlcNAc6S | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Fuc(a1-2)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)Glc6S | Fuc(a1-2)Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)GalNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)GlcNAc6S | Fuc(a1-2)Gal(b1-4)[Fuc(a1-2)]Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GalNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GalNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GalNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNA | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)]Man(a1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-3)GalNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)GalNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]GalNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Fuc(a1-2)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)GalNAc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-3)]GalNAc | Fuc(a1-2)Gal3S | Fuc(a1-2)Gal3S(b1-4)GlcNAc | Fuc(a1-2)Gal6S(b1-3)GlcNAc | Fuc(a1-2)Gal6S(b1-3)GlcNAc6S | Fuc(a1-2)Gal6S(b1-4)Glc | Fuc(a1-2)Gal6S(b1-4)Glc6S | Fuc(a1-2)Gal6S(b1-4)GlcNAc | Fuc(a1-2)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal | Fuc(a1-2)[Gal(a1-3)]Gal(a1-4)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-3)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)GlcNAc(b1-6)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-3)[Fuc(a1-4)]GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-6)]GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)GlcNAc(b1-6)GalNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)[Gal(a1-3)]Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)GalNAc | Fuc(a1-2)[Gal(a1-4)]Gal(b1-4)GalNAc | Fuc(a1-2)[Gal(a1-4)]Gal(b1-4)GlcNAc | Fuc(a1-2)[Gal(b1-3)]Gal | Fuc(a1-2)[Gal(b1-3)]Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-2)[GalN(a1-3)]Gal | Fuc(a1-2)[GalNAc(a1-3)]Gal | Fuc(a1-2)[GalNAc(a1-3)]Gal(a1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(a1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)GlcNAc(b1-6)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)[Fuc(a1-4)]GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-6)]GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)GlcNAc(b1-6)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)[Fuc(a1-2)]Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)[GalNAc(a1-3)]Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)GalNAc | Fuc(a1-2)[GalNAc(a1-3)]GalNAc | Fuc(a1-2)[GalNAc(a1-4)]Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(b1-3)]Gal | Fuc(a1-2)[GalNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-2)[GalNAc(b1-3)]Gal(b1-4)GlcNAc | Fuc(a1-2)[GalNAc(b1-3)]Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-2)[GalNGc(a1-3)]Gal | Fuc(a1-2)[Neu5Ac(a2-3)]Gal(b1-4)GlcNAc(b1-3)GalNAc | Fuc(a1-2)[Neu5Ac(a2-6)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-2)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Fuc(a1-2)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-2)[Neu5Ac(b2-6)]Gal(b1-4)GlcNAc | Fuc(a1-2)[Neu5Ac(a2-3)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-3)GlcNAc | Fuc(a1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-3)GlcNAc6S | Fuc(a1-3)Xluf(b1-3)Fuc | Fuc(a1-3)Xulf(b1-3)Fuc | Fuc(a1-3)[3dGal(b1-4)]Glc | Fuc(a1-3)[3dGal(b1-4)]GlcNAc(b1-3)Gal | Fuc(a1-3)[Fuc(a1-6)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]Glc | Fuc(a1-3)[Gal(b1-4)]Glc6S | Fuc(a1-3)[Gal(b1-4)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-3)GalNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)GalNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[Fuc(a1-2)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)GalNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)Glc | Fuc(a1-3)[Gal(b1-4)]GlcNAc6S | Fuc(a1-3)[Gal3S(b1-4)]Glc | Fuc(a1-3)[Gal3S(b1-4)]Glc6S | Fuc(a1-3)[Gal3S(b1-4)]GlcNAc | Fuc(a1-3)[Gal3S(b1-4)]GlcNAc(b1-3)Gal | Fuc(a1-3)[Gal3S(b1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[Gal3S(b1-4)]GlcNAc6S | Fuc(a1-3)[Gal3S(b1-4)]GlcNAc6S(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[Gal6S(b1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-3)[GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Fuc(a1-3)[GalNAc(b1-4)]GlcNAc6S | Fuc(a1-3)[GalNAc3S(b1-4)]GlcNAc | Fuc(a1-3)[GlcNAc(b1-4)]GlcNAc | Fuc(a1-3)[Man(b1-4)]GlcNAc(b1-4)GlcNAc | Fuc(a1-4)Fuc(a1-2)Glc(b1-3)GlcNAc(a1-3)GlcA(b1-4)[Glc(a1-3)]Fuc | Fuc(a1-4)GlcNAc | Fuc(a1-6)GlcNAc | Fuc(b1-2)Gal(b1-3)GlcNAc | Fuc(b1-2)Gal(b1-4)GlcNAc | Fuc(b1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Fuc(b1-3)GlcNAc | Fuc(b1-3)[Gal(b1-4)]GlcNAc | Fuc1P(b1-4)Glc(a1-3)Fuc(a1-3)GlcNAc(b1-2)[6dTal(a1-3)]Fuc1P | Fuc3Ac4Ac(a1-2)Gal(b1-3)GalNAc(a1-3)GalNAc(a1-2)Fuc3Ac4Ac | Fuc3NBut4Ac(b1-6)Glc3Ac(a1-4)GlcA(b1-3)GlcNAc(a1-2)Fuc3NBut4Ac | FucNAc | FucNAc(a1-3)D-FucNAc | FucNAc(a1-3)GalNAc(a1-4)Gal | FucNAc(a1-3)GlcNAc(a1-3)GlcA(b1-4)[Gal(a1-3)]FucNAc | FucNAc(a1-3)GlcNAc(b1-3)Glc6PGro(b1-4)[Glc(a1-6)Gal2Ac(a1-3)]FucNAc | FucNAc(a1-3)GlcNAc(b1-4)[GlcA(a1-3)]FucNAc | FucNAc(a1-3)GlcNAc(b1-6)Glc6PGro(b1-4)[Glc(a1-6)Gal2Ac(a1-3)]FucNAc | FucNAc(a1-3)GlcNAc6PEtN(a1-3)GlcA(b1-4)[Gal(a1-3)]FucNAc | FucNAc(b1-3)GalNAc(a1-4)Gal | Fucf2Ac(b1-3)DDManHep(b1-3)Fucf2Ac | Fucf2Ac(b1-3)LDManHep(b1-3)Fucf2Ac | Gal | Gal(a1-2)Gal | Gal(a1-2)Gal(a1-2)[Gal(b1-4)]Glc(a1-3)Glc | Gal(a1-3)FucNAc(a1-3)GalNAc(b1-4)[Glc(a1-2)Glc(a1-3)]ManNAcA | Gal(a1-3)Gal | Gal(a1-3)Gal(b1-3)Glc | Gal(a1-3)Gal(b1-3)GlcNAc | Gal(a1-3)Gal(b1-3)GlcNAc(b1-2)Man | Gal(a1-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-3)GlcNAc(b1-3)GalNAc | Gal(a1-3)Gal(b1-3)GlcNAc(b1-6)GalNAc | Gal(a1-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal | Gal(a1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-2)Man | Gal(a1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)GalNAc | Gal(a1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GalNAc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-3)GalNAc | Gal(a1-3)GalNAc6S | Gal(a1-3)GalfNAc | Gal(a1-3)GlcNAc | Gal(a1-3)ManNAc(b1-6)Galf(b1-3)GlcNAc(b1-2)[Galf(a1-4)]Gal | Gal(a1-3)ManNAc(b1-6)Galf(b1-3)GlcNAc(b1-2)[Galf2Ac(a1-4)]Gal | Gal(a1-3)[Gal(a1-4)]Gal(b1-4)GlcNAc | Gal(a1-3)[GlcA(b1-4)]Fuc(a1-4)GlcNAc(b1-3)Fuc(a1-3)GlcNAc(b1-3)Gal | Gal(a1-3)[Neu5Ac(a2-6)]GalNAc | Gal(a1-3)[Neu5Ac(b2-6)]GalNAc | Gal(a1-4)Gal | Gal(a1-4)Gal(b1-3)GlcNAc | Gal(a1-4)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(a1-4)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-4)Gal(b1-4)Gal | Gal(a1-4)Gal(b1-4)GalNAc | Gal(a1-4)Gal(b1-4)Glc | Gal(a1-4)Gal(b1-4)GlcNAc | Gal(a1-4)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(a1-4)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-4)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(a1-4)GalNAc | Gal(a1-4)GalNAc(a1-3)FucNAc(a1-3)GlcA6PEtN(b1-3)Gal | Gal(a1-4)GalNAc(a1-3)FucNAc(a1-3)GlcNAc6PEtN(b1-3)Gal | Gal(a1-4)GalNAc(b1-3)GalNAc(b1-3)[Dhp(a2-4)Man(b1-4)]Gal | Gal(a1-4)GalNAc(b1-3)GalNAc(b1-3)[Dhpa(2-4)Man(b1-4)]Gal | Gal(a1-4)Glc | Gal(a1-4)GlcA(b1-4)Glc(b1-4)Glc | Gal(a1-4)GlcA(b1-6)Gal(b1-4)Gal(b1-4)GlcNAc(b1-6)Gal | Gal(a1-4)GlcNAc | Gal(a1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(a1-6)Glc | Gal(a1-6)Glc(b1-3)GalNAc(b1-3)[GalNAc4PGro(b1-4)]Gal | Gal(a1-6)Glc(b1-3)GalNAc(b1-4)[GalNAc(b1-3)]Gal | Gal(a1-6)Glc(b1-3)GalNAc(b1-4)[Rha(a1-4)GalNAc(b1-3)]Gal | Gal(a1-6)Glc(b1-3)[Glc(b1-6)]GalNAc(b1-4)[Glc(b1-2)][GlcA3Ac4Ac(b1-3)]Gal | Gal(a1-6)Man(a1-2)Man(a1-3)GalNAc(b1-3)[GlcA(b1-4)]Gal | Gal(a1-6)Man(a1-2)Man(a1-3)GalNAc(b1-3)[Ribf(b1-4)GlcA(b1-4)]Gal | Gal(b1-2)Gal | Gal(b1-2)Gal(a1-3)GlcNAc | Gal(b1-2)Gal(a1-4)GlcNAc | Gal(b1-2)Gal(b1-4)GlcNAc | Gal(b1-3)Ara | Gal(b1-3)Fuc | Gal(b1-3)FucNAc(a1-3)GlcNAc(b1-2)Gal | Gal(b1-3)Gal | Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-4)Glc | Gal(b1-3)Gal(b1-3)Gal(b1-3)Gal(b1-4)Glc | Gal(b1-3)Gal(b1-3)Gal(b1-4)Glc | Gal(b1-3)Gal(b1-4)Glc | Gal(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GalNAc | Gal(b1-3)GalNAc(a1-3)GlcNAc(b1-6)Glc(a1-4)[GlcNAc(b1-3)]Gal | Gal(b1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)Glc | Gal(b1-3)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)GlcNAc | Gal(b1-3)GalNAc(a1-4)Gal(b1-3)GalNAc(b1-4)[GlcNAc(b1-2)Glc(b1-2)][GlcNAc(b1-3)]Gal | Gal(b1-3)GalNAc(b1-3)Gal | Gal(b1-3)GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)GalNAc(b1-3)GalNAc(b1-2)Gal(a1-3)[Fuc(a1-2)]Gal | Gal(b1-3)GalNAc(b1-4)Gal(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)Gal2S(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)Gal3S(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)Gal3S(b1-4)GlcNAc(b1-3)GalNAc | Gal(b1-3)GalNAc(b1-4)[Kdn(a2-3)]Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Gal(b1-3)GalNAc(b1-4)[Neu5Gc(a2-3)]Gal(b1-4)Glc | Gal(b1-3)GalNAc6S | Gal(b1-3)GalNAc6S(b1-4)Gal(b1-4)Glc | Gal(b1-3)GalNAc6S(b1-4)Gal3S(b1-4)Glc | Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Gal(b1-3)GalNGc | Gal(b1-3)GalfNAc | Gal(b1-3)GlcNAc | Gal(b1-3)GlcNAc(a1-3)Gal(b1-3)GlcNAc | Gal(b1-3)GlcNAc(a1-3)Gal(b1-4)GlcNAc | Gal(b1-3)GlcNAc(a1-3)GalNAc | Gal(b1-3)GlcNAc(a1-3)Glc6PEtN(b1-3)GlcNAc(b1-2)[Rib5P-ol(5-3)]Gal | Gal(b1-3)GlcNAc(a1-4)Rha(a1-3)GlcNAc(b1-2)Gal | Gal(b1-3)GlcNAc(a1-6)Gal(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-3)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal | Gal(b1-3)GlcNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-3)GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)GalNAc | Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)]GlcNAc(b1-6)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)]Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal | Gal(b1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal | Gal(b1-3)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)Glc | Gal(b1-3)GlcNAc(b1-6)Gal | Gal(b1-3)GlcNAc(b1-6)Gal(b1-4)GlcNAc | Gal(b1-3)GlcNAc(b1-6)GalNAc | Gal(b1-3)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Gal(b1-3)GlcNAc6Bz | Gal(b1-3)GlcNAc6S | Gal(b1-3)GlcNFm(b1-3)Gal(b1-3)GlcNAc | Gal(b1-3)GlcNFm(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)GlcNFo(b1-3)Gal(b1-3)GlcNAc | Gal(b1-3)GlcNFo(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)Man | Gal(b1-3)[Fuc(a1-4)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc.1 | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-6)GalNAc | Gal(b1-3)[Fuc(b1-4)]GlcNAc | Gal(b1-3)[Gal(b1-4)]GlcNAc | Gal(b1-3)[Gal(b1-6)]GalNAc | Gal(b1-3)[GlcNAc(b1-6)]GalNAc | Gal(b1-3)[GlcNAc(b1-6)]GlcNAc | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(a1-4)Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-4)Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-6)Gal(b1-4)Glc | Gal(b1-3)[Neu5Ac(b2-6)]GalNAc | Gal(b1-3)[Neu5Ac(b2-6)]GlcNAc | Gal(b1-4)Fuc | Gal(b1-4)Gal | Gal(b1-4)Gal(a1-3)Gal | Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal | Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal | Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal | Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal | Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal(b1-4)Gal(a1-3)Gal | Gal(b1-4)Gal(b1-4)Glc | Gal(b1-4)Gal(b1-4)GlcNAc | Gal(b1-4)GalNAc | Gal(b1-4)GalNAc(a1-3)[Fuc(a1-2)]Gal(b1-4)GlcNAc | Gal(b1-4)GalNAc(b1-3)[Fuc(a1-2)]Gal(b1-4)GlcNAc | Gal(b1-4)Glc | Gal(b1-4)Glc(a1-4)GlcA(a1-3)GalNAc(b1-3)[Galf(a1-2)Rha(a1-4)]Gal | Gal(b1-4)Glc(b1-6)[Gal(b1-4)]GlcNAc | Gal(b1-4)Glc6S | Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(a1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(a1-6)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Gal(b1-4)GlcNAc(b1-2)GalNAc | Gal(b1-4)GlcNAc(b1-2)Man | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][Gal(b1-4)GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Gal(b1-4)Fuc(a1-6)][Fuc(a1-3)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc6S(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-3)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-4)][Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Fuc | Gal(b1-4)GlcNAc(b1-3)Gal | Gal(b1-4)GlcNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(a1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(a1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)Gal(b1-6)[Gal(b1-3)]GalNAc | Gal(b1-4)GlcNAc(b1-3)GalNAc | Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-3)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Kdn(a2-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]GalNAc | Gal(b1-4)GlcNAc(b1-4)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[Man(a1-3)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-4)[GlcNAc(b1-2)]Man(a1-3)[Man(a1-6)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-6)Gal | Gal(b1-4)GlcNAc(b1-6)Gal(b1-4)GlcNAc | Gal(b1-4)GlcNAc(b1-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-6)Gal(b1-4)GlcNAc(b1-6)GalNAc | Gal(b1-4)GlcNAc(b1-6)GalNAc | Gal(b1-4)GlcNAc(b1-6)Man | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GlcNAc | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]GalNAc | Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)Glc | Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Man | Gal(b1-4)GlcNAc6P | Gal(b1-4)GlcNAc6S | Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc6S(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal(b1-4)GlcNAc6S(b1-3)Gal(b1-4)GlcNAc6S | Gal(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S | Gal(b1-4)Man | Gal(b1-4)Man(b1-4)Gal(a1-3)GlcNAc(b1-2)Gal | Gal(b1-4)ManNAc | Gal(b1-4)[Glc(b1-6)]GlcNAc(b1-3)Gal | Gal(b1-4)[Neu5Ac(a2-6)]GlcNAc | Gal(b1-6)Gal | Gal(b1-6)Gal(b1-4)Glc | Gal(b1-6)GalNAc | Gal(b1-6)Galf(b1-3)GalNAc(b1-3)[Glc4Lac(b1-6)Glc(a1-4)]Gal | Gal(b1-6)GlcNAc | Gal(b1-6)Man | Gal2S(b1-4)Gal2S6S | Gal2S(b1-4)Gal2S6S(a1-3)Gal2S | Gal2S(b1-4)Gal2S6S(a1-3)Gal2S(b1-4)Gal2S6S | Gal2S(b1-4)Gal2S6S(a1-3)Gal2S(b1-4)Gal2S6S(a1-3)Gal2S | Gal2S(b1-4)Gal2S6S(a1-3)Gal2S(b1-4)Gal2S6S(a1-3)Gal2S(b1-4)Gal2S6S | Gal3Pyr | Gal3Pyr4Pyr(b1-4)Man(b1-4)Gal(a1-3)GlcNAc(b1-2)Gal3Pyr4Pyr | Gal3S | Gal3S(b1-3)GalNAc | Gal3S(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Gal3S(b1-3)GalNAc(b1-4)Gal3S(b1-4)Glc | Gal3S(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Gal3S(b1-3)GalNAc6S(b1-4)Gal(b1-4)Glc | Gal3S(b1-3)GalNAc6S(b1-4)Gal3S(b1-4)Glc | Gal3S(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Gal3S(b1-3)GlcNAc | Gal3S(b1-3)[Fuc(a1-4)]GlcNAc | Gal3S(b1-3)[Fuc(a1-4)]GalNAc | Gal3S(b1-3)[Fuc(a1-4)]GlcNAc.1 | Gal3S(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal3S(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal(b1-4)Glc | Gal3S(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal3S(b1-4)Glc | Gal3S(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Gal3S(b1-4)Glc | Gal3S(b1-4)Glc6S | Gal3S(b1-4)GlcNAc | Gal3S(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal3S(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal3S(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal3S(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal3S(b1-4)GlcNAc(b1-3)Gal | Gal3S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal3S(b1-4)GlcNAc(b1-3)GalNAc | Gal3S(b1-4)GlcNAc6S | Gal3S(b1-4)[Fuc(a1-3)]Glc6S | Gal3S(b1-4)[Fuc(a1-3)]GlcNAc | Gal3S(b1-4)[Fuc(a1-3)]GlcNAc6S | Gal3S4S(b1-4)GlcNAc | Gal3S6S(b1-4)GlcNAc | Gal3S6S(b1-4)GlcNAc6S | Gal4S | Gal4S(b1-4)Gal(a1-3)Gal4S | Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S | Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S | Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S(b1-4)Gal(a1-3)Gal4S | Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S | Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S | Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S | Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S(a1-3)Gal4S(b1-4)Gal2S | Gal4S(b1-4)GlcNAc | Gal4S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal4S6S(b1-4)GlcNAc | Gal6Bz(a1-4)GlcNAc6Bz | Gal6Bz(b1-3)GlcNAc | Gal6Bz(b1-3)GlcNAc6Bz | Gal6Bz(b1-4)GlcNAc | Gal6Bz(b1-4)GlcNAc6Bz | Gal6P(b1-4)GlcNAc | Gal6S | Gal6S(a1-3)GalNAc | Gal6S(b1-3)GalNAc | Gal6S(b1-3)GlcNAc | Gal6S(b1-3)GlcNAc6S | Gal6S(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Gal6S(b1-4)Gal(a1-3)Gal6S | Gal6S(b1-4)Gal(a1-3)Gal6S(b1-4)Gal(a1-3)Gal6S | Gal6S(b1-4)Glc | Gal6S(b1-4)Glc6S | Gal6S(b1-4)GlcNAc | Gal6S(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal6S(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal6S(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal6S(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Gal6S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Gal6S(b1-4)GlcNAc6S | Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S | GalA(a1-2)Rha(a1-2)Ribf(b1-4)Gal(b1-3)GlcNAc(b1-4)GalA | GalA(a1-3)FucNAc(a1-3)GlcNAc(b1-3)FucNAc(a1-4)[GlcNAc(b1-2)]GalA | GalA6Lys(a1-4)Gal(a1-3)GalA4Ac6Ser(a1-3)GlcNAc(b1-4)GalA6Lys | GalA6N(a1-4)GalNAc(a1-4)Gal(a1-3)GalNAc(b1-3)[Qui3NFo(a1-4)]GalA6N | GalN(a1-6)ManN(a1-3)Fuc(a1-3)[Glc(b1-4)]Gal(b1-4)[Fuc(a1-3)]GalN | GalN6Ac(a1-3)D-Fuc2Ac(a1-3)GlcNAc6PEtN(b1-3)Gal(a1-4)GalN6Ac | GalNAc | GalNAc(a1-2)Glc(a1-4)IdoA(a1-3)GalNAc(b1-4)[GlcNAc(b1-3)]GalNAc | GalNAc(a1-3)Gal | GalNAc(a1-3)Gal(b1-4)Glc | GalNAc(a1-3)Gal(b1-4)GlcNAc | GalNAc(a1-3)Gal(b1-4)GlcNAc(a1-3)Gal(b1-4)GlcNAc | GalNAc(a1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | GalNAc(a1-3)GalA(a1-3)GlcNAc(a1-4)[AltA(a1-3)]GalNAc | GalNAc(a1-3)GalNAc | GalNAc(a1-3)GalNAc(a1-3)Gal(a1-4)Gal(b1-4)GlcNAc | GalNAc(a1-3)GalNAc(b1-3)Gal | GalNAc(a1-3)GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | GalNAc(a1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | GalNAc(a1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc | GalNAc(a1-3)GalNAc(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | GalNAc(a1-3)GalfNAc | GalNAc(a1-4)Gal | GalNAc(a1-4)Gal(b1-4)GlcNAc | GalNAc(a1-4)GalNAc | GalNAc(a1-4)Glc(a1-4)IdoA(a1-3)GalNAc(b1-4)[GlcNAc(b1-3)]GalNAc | GalNAc(a1-4)GlcA(b1-3)GlcNAc(b1-2)GalA(b1-3)GalNAc | GalNAc(a1-4)GlcNAc3NAcA(b1-3)D-FucNAc(a1-3)QuiNAc(b1-3)Rha(a1-3)[Glc(a1-6)]Glc(b1-3)[Glc(a1-4)]GalN2Ala6P(a1-3)LDManHep2P4P7Cm(a1-3)LDManHep(a1-5)Kdo | GalNAc(a1-6)Man2Ac3Ac4Ac(a1-3)Fuc(a1-3)[Glc(b1-4)]Gal(b1-4)[Fuc(a1-3)]GalNAc | GalNAc(b1-2)Gal(a1-3)[Fuc(a1-2)]Gal(b1-3)GalNAc(b1-3)GalNAc | GalNAc(b1-3)Gal | GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc | GalNAc(b1-3)Gal(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GalNAc(b1-3)Gal(a1-6)Gal(b1-4)Glc | GalNAc(b1-3)Gal(b1-4)Glc | GalNAc(b1-3)Gal(b1-4)GlcNAc | GalNAc(b1-3)GalNAc | GalNAc(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | GalNAc(b1-3)GlcNAc | GalNAc(b1-3)GlcNAc(b1-2)[Glc3Lac(a1-3)]Rha(a1-2)Ribf(b1-4)GalNAc | GalNAc(b1-3)Rha(b1-4)GlcNAc6Ac(b1-4)[GalA3Ac6Thr(a1-3)]GalNAc | GalNAc(b1-3)Rha(b1-4)GlcNAc6Ac(b1-4)[GalA6Thr3Ac(a1-3)]GalNAc | GalNAc(b1-3)[Gal(b1-4)]Gal(b1-4)[GlcNAc(b1-2)]Glc(b1-3)GalNAc | GalNAc(b1-4)Gal | GalNAc(b1-4)Gal(b1-4)Glc | GalNAc(b1-4)Gal3S(b1-4)Glc | GalNAc(b1-4)Gal3S(b1-4)GlcNAc(b1-3)GalNAc | GalNAc(b1-4)GalNAc | GalNAc(b1-4)GalNAc(a1-3)GlcNAc(a1-2)Rib-ol | GalNAc(b1-4)GalNAc(b1-4)GalNAc | GalNAc(b1-4)GalNAc3PGro(b1-4)[Gal(a1-6)]Glc(b1-3)GalNAc(b1-6)GalNAc | GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA | GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA | GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA | GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA | GalNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[GalNAc(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GalNAc(b1-4)GlcNAc(b1-3)Gal(b1-6)[Gal(b1-3)]GalNAc | GalNAc(b1-4)GlcNAc(b1-3)GalNAc | GalNAc(b1-4)GlcNAc(b1-3)GalNAc(b1-4)GlcNAc | GalNAc(b1-4)GlcNAc(b1-6)GalNAc | GalNAc(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | GalNAc(b1-4)GlcNAc6Bz | GalNAc(b1-4)GlcNAc6S | GalNAc(b1-4)Man | GalNAc(b1-4)[Neu5Ac(a2-6)]Gal(b1-4)Glc | GalNAc(b1-6)GalNAc | GalNAc2S4S | GalNAc2S4S6S | GalNAc2S6S | GalNAc3Ac4Ac(a1-4)GalNAc3Ac(a1-4)GalNAc3Ac(a1-4)GalNAc3Ac | GalNAc3PGro(b1-3)Gal(a1-4)Gal(b1-3)GalNAc(b1-4)GalNAc3PGro | GalNAc3PGroAN(b1-4)Gal(b1-3)FucNAc4N(b1-4)GalNAc3PGroAN | GalNAc3S | GalNAc3S(b1-4)Gal3S(b1-4)Glc | GalNAc3S(b1-4)GlcNAc | GalNAc3S(b1-4)GlcNAc3Ac | GalNAc3S(b1-4)GlcNAc3S | GalNAc3S(b1-4)[Fuc(a1-3)]GlcNAc | GalNAc3S4S(b1-4)GlcNAc | GalNAc3S6S(b1-4)GlcNAc | GalNAc4S | GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | GalNAc4S(b1-4)GlcNAc | GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | GalNAc4S6S | GalNAc4S6S(b1-4)GlcNAc | GalNAc4S6S(b1-4)GlcNAc3Ac | GalNAc6S | GalNAc6S(b1-4)Gal(b1-4)Glc | GalNAc6S(b1-4)Gal3S(b1-4)Glc | GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GalNAc6S(b1-4)GlcNAc | GalNAc6S(b1-4)GlcNAc3Ac | GalNAc6S(b1-4)GlcNAc3S | GalNAc6S(b1-4)GlcNAc6S | GalNAcA3Ac6N(a1-4)GalNAcA(a1-3)GlcNAc | GalNGc | GalNGc(a1-3)GalNAc | Galf(b1-3)Galf(b1-5)[Glc(a1-2)]Galf(b1-5)[Glc(a1-2)]Galf | Galf(b1-3)GlcNAc(a1-8)Pse5Ac7But(a2-6)Gal(a1-6)Glc(a1-2)Galf | Galf2Ac(b1-3)Gal(a1-3)Galf2Ac | GalfNAc(b1-4)GlcNAc | Glc | Glc(a1-1)Glc | Glc(a1-2)Gal | Glc(a1-2)Gal(a1-3)Glc | Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc(a1-2)Glc | Glc(a1-2)Glc(a1-2)[GlcNAc(a1-3)]Gal(a1-3)Glc | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Glc(a1-2)Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Glc(a1-2)Glc(a1-3)[FucNAc(a1-3)GalNAc(b1-4)]ManNAcA | Glc(a1-3)GalNAc(a1-3)GalNAc(b1-4)[Rha(a1-3)][Glc(a1-6)]Glc | Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc(a1-3)Glc | Glc(a1-3)Glc(a1-3)Man | Glc(a1-3)Glc(a1-3)Man(a1-2)Man | Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man | Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-3)Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Glc(a1-4)Gal | Glc(a1-4)Gal(a1-3)GlcNAc(b1-4)[Col(a1-3)][Col(a1-6)]Glc | Glc(a1-4)Gal(a1-4)GlcA(b1-4)Glc | Glc(a1-4)GalNAc(b1-4)Man | Glc(a1-4)GalNAc(b1-4)[Man(a1-2)Man(a1-6)]Man | Glc(a1-4)GalNAc(b1-4)[Man6PEtN(a1-2)Man(a1-6)]Man | Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-4)Glc(a1-6)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-4)Glc(a1-6)Glc | Glc(a1-4)GlcA(b1-3)GlcNAc(a1-3)Rha(a1-4)Glc | Glc(a1-6)Glc | Glc(a1-6)Glc(a1-4)Glc | Glc(a1-6)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-6)Glc(a1-4)Glc(a1-4)Glc(a1-6)Glc(a1-4)Glc(a1-4)Glc | Glc(a1-6)Glc(a1-4)[GlcNAc(b1-3)]Gal(a1-3)GlcNAc(b1-2)Glc | Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc(a1-6)Glc | Glc(a1-6)Glc(b1-3)[Rha(a1-6)Glc(a1-4)]GalN(a1-3)LDManHep6P(a1-3)LDManHep2P4P(a1-5)[Kdo(a2-4)]Kdo(a2-6)GlcN4P(b1-6)GlcN1P | Glc(b1-2)Fuc3NHb(b1-6)GlcNAc(a1-4)GalNAc(a1-3)[Glc(a1-6)]GlcNAc(b1-2)[Glc(a1-4)]Glc | Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc(b1-2)Glc | Glc(b1-3)6dTal2Ac(a1-3)GlcNAc(b1-2)Glc | Glc(b1-3)D-FucNAc | Glc(b1-3)FucNAc | Glc(b1-3)Gal(a1-4)GalNAc(b1-4)GlcA6Ser(b1-3)GalNAc(b1-4)Glc | Glc(b1-3)Gal(b1-3)GalNAc(b1-4)Rib5P-ol(5-4)[Glc(a1-3)]Glc | Glc(b1-3)Gal(b1-4)Man | Glc(b1-3)GalNAc | Glc(b1-3)GalNAc(b1-3)[Gal(b1-4)]Gal(b1-4)[GlcNAc(b1-2)]Glc | Glc(b1-3)GalNAc(b1-4)GalNAc(b1-4)Gal(b1-4)Glc | Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-3)Glc(b1-4)Glc | Glc(b1-3)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-3)Glc(b1-4)Glc(b1-6)Glc(b1-4)Glc | Glc(b1-3)GlcA | Glc(b1-3)GlcA(b1-4)Glc | Glc(b1-3)GlcNAc | Glc(b1-3)GlcNAc(b1-3)[Rib1Ac5P-ol(5-6)]Gal(b1-4)[GlcNAc(b1-2)]Glc | Glc(b1-3)L-6dTal2Ac(a1-3)GlcNAc(b1-2)Glc | Glc(b1-3)[Glc(b1-6)]Glc(a1-3)Glc(b1-3)[Glc(b1-6)]Glc | Glc(b1-3)[Glc(b1-6)]Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc(b1-3)Glc | Glc(b1-4)Gal(b1-4)Glc | Glc(b1-4)GalNAc | Glc(b1-4)Glc | Glc(b1-4)Glc(a1-2)Glc | Glc(b1-4)Glc(a1-4)Gal(a1-4)GlcA | Glc(b1-4)Glc(b1-3)Glc | Glc(b1-4)Glc(b1-3)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-3)GlcA(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-3)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-3)Glc(b1-4)Glc(b1-4)Glc(b1-3)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-3)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc(b1-4)Glc | Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Glc(b1-6)Gal(a1-6)GalNAc(a1-4)[Glc(b1-3)]GalNAc(a1-3)GalNAc(a1-2)Glc | Glc(b1-6)Glc | Glc(b1-6)Glc(b1-3)Glc | Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc(b1-6)Glc | Glc(b1-6)GlcNAc(a1-3)FucNAc(a1-3)GlcNAc(b1-2)Glc | Glc-ol | Glc3Me6Me | Glc3Me6Me(b1-4)Rha2Me3Me | Glc4Lac(b1-4)GalNAc(a1-3)GalNAc(b1-6)Glc4Lac | Glc4S(a1-4)Glc3S | Glc6Ac(a1-3)GlcA4Ac(b1-3)GlcNAc(a1-3)GlcA4Ac(b1-4)Glc6Ac | Glc6P | Glc6S | Glc6S(a1-4)Glc6S | GlcA | GlcA(a1-2)Rha(a1-2)Rha(a1-2)Gal(a1-3)GalNAc(b1-3)[Rha(a1-4)]GlcA | GlcA(a1-3)Gal | GlcA(a1-3)Gal(a1-3)ManNAc(b1-4)Glc(b1-4)Glc | GlcA(a1-3)Gal(a1-3)ManNAc6Ac(b1-4)Glc(b1-4)Glc | GlcA(b1-2)[Gal(a1-3)]Man6Ac(b1-4)Man(b1-3)GlcNAc(b1-4)GlcA | GlcA(b1-3)Gal | GlcA(b1-3)Gal(b1-3)GlcNAc | GlcA(b1-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | GlcA(b1-3)Gal(b1-4)Glc | GlcA(b1-3)Gal(b1-4)GlcNAc | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | GlcA(b1-3)GalA6Thr(b1-3)[Glc(a1-6)]GlcNAc(b1-4)[GlcNAc(b1-2)]GlcA | GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-3)[GlcNAc(b1-4)Glc(b1-4)]GalNAc(b1-4)GlcA | GlcA(b1-3)GalNAc(b1-4)GlcA | GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc(b1-4)GlcA(b1-3)GalNAc | GlcA(b1-3)GalNAc(b1-6)[GalA6Lys(a1-4)]GalNAc(b1-4)[Glc(a1-2)]GlcA | GlcA(b1-3)GalNAc4S | GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | GlcA(b1-3)GalNAc4S6S | GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | GlcA(b1-3)GlcN(a1-4)GlcA(b1-3)Man | GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)Man | GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)Man | GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)Man | GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-3)Man | GlcA(b1-3)GlcNAc(a1-4)GlcA(b1-4)[GlcNAc(b1-2)]GlcA | GlcA(b1-3)GlcNAc(b1-2)Qui4NAlaBut(b1-3)Gal(a1-4)GlcA | GlcA(b1-3)GlcNAc(b1-2)Qui4NButAla(b1-3)Gal(a1-4)GlcA | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | GlcA(b1-4)FucNAc | GlcA(b1-4)GalNAc(a1-4)GlcA(b1-4)GalNAc4S | GlcA(b1-4)GalNAc4S(a1-4)GlcA(b1-4)GalNAc4S | GlcA(b1-4)Glc | GlcA(b1-4)Glc(b1-3)GlcA | GlcA(b1-4)Glc(b1-3)GlcA(b1-4)Glc | GlcA(b1-4)Glc(b1-4)Glc(a1-4)Gal | GlcA(b1-4)GlcNAc(a1-4)GlcA | GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)2,5-Anhydro-Man | GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)2,5-Anhydro-Man | GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA(b1-4)Man | GlcA(b1-4)GlcNAc(a1-4)GlcA2S(b1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)GlcA2S(b1-4)GlcNAc6S | GlcA(b1-4)GlcNAc(a1-4)IdoA(a1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)IdoA(a1-4)GlcNAc6S | GlcA(b1-4)GlcNAc(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc6S | GlcA(b1-4)GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNAc(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNAc(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc(b1-3)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNAc6S | GlcA(b1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc6S | GlcA(b1-4)GlcNAc6S(a1-4)GlcA2S(b1-4)GlcNAc6S | GlcA(b1-4)GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc6S | GlcA(b1-4)GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc6S | GlcA(b1-4)GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNS(a1-4)GlcA2S(b1-4)GlcNS | GlcA(b1-4)GlcNS(a1-4)GlcA2S(b1-4)GlcNS6S | GlcA(b1-4)GlcNS(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS | GlcA(b1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNS3S(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNS3S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS3S6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNAc | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS3S6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)GlcA2S(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNAc6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS3S6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS3S6S(a1-4)GlcA(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS3S6S(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS3S6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS | GlcA(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | GlcA(b1-6)Gal | GlcA2S(b1-3)GalNAc4S(b1-4)GlcA2S(b1-3)GalNAc4S | GlcA2S(b1-3)GalNAc6S | GlcA3Ac(b1-3)GlcNAc(a1-2)Fuc3NBut(b1-6)Glc4Ac(a1-4)[Glc(a1-2)]GlcA3Ac | GlcA3S(b1-3)Gal | GlcA3S(b1-3)Gal(b1-4)Glc | GlcA3S(b1-3)Gal(b1-4)GlcNAc | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | GlcA6Ala(b1-3)GalNAc(b1-4)Glc(b1-3)Gal(a1-4)GalNAc(b1-4)GlcA6Ala | GlcA6CetLys(b1-6)GalNAc(a1-6)GlcNAc(b1-3)GlcNAc(b1-4)GlcA6CetLys | GlcA6Thr3Ac(b1-6)Gal(b1-6)Glc(b1-3)GalNAc6Ac(b1-4)GlcA6Thr3Ac | GlcA6ThrAc(b1-6)Gal(b1-6)Glc(b1-3)GalNAc6Ac(b1-4)GlcA6ThrAc | GlcN | GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN | GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN | GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN(b1-4)GlcN | GlcN2S | GlcN2S6S | GlcN6S | GlcNAc | GlcNAc(a1-2)Glc(a1-2)Glc(a1-3)[Gal(a1-6)]Glc | GlcNAc(a1-2)LDManHep(a1-3)LDManHep | GlcNAc(a1-2)LDManHep(a1-3)LDManHep(a1-5)Kdo | GlcNAc(a1-3)Gal(b1-3)GalNAc(a1-6)GlcNAc | GlcNAc(a1-3)Gal(b1-4)GlcNAc | GlcNAc(a1-3)GalNAc | GlcNAc(a1-3)GlcNAc(b1-6)GlcNAc6P(a1-3)GlcNAc6PPenn | GlcNAc(a1-3)GlcNAc6PPenn | GlcNAc(a1-3)Qui4NBut(b1-4)[Gal(a1-3)]Man(a1-4)Rha(a1-3)GlcNAc | GlcNAc(a1-3)[Fuc3Ac4Ac(a1-4)]GlcNAc(a1-4)GlcA(a1-3)Fuc(a1-3)GlcNAc | GlcNAc(a1-4)Gal | GlcNAc(a1-4)Gal(b1-3)GalNAc | GlcNAc(a1-4)Gal(b1-3)GlcNAc | GlcNAc(a1-4)Gal(b1-3)[Fuc(a1-2)]Gal | GlcNAc(a1-4)Gal(b1-4)GalNAc | GlcNAc(a1-4)Gal(b1-4)GlcNAc | GlcNAc(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | GlcNAc(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | GlcNAc(a1-4)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(a1-4)GlcA(b1-4)GlcNAc(a1-4)GlcA | GlcNAc(a1-4)GlcNAc | GlcNAc(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA | GlcNAc(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA2S | GlcNAc(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA | GlcNAc(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNAc(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA | GlcNAc(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA2S | GlcNAc(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA | GlcNAc(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA | GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA2S | GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA | GlcNAc(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNAc(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA | GlcNAc(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA2S | GlcNAc(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA | GlcNAc(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNAc(a1-6)Gal(b1-4)GlcNAc | GlcNAc(b1-2)Fuc | GlcNAc(b1-2)Gal | GlcNAc(b1-2)Gal(b1-3)GalNAc | GlcNAc(b1-2)Gal(b1-3)GlcNAc | GlcNAc(b1-2)Man | GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)Man(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-3)[Xyl(b1-2)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)Man(a1-6)[Xyl(b1-2)][Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)Rha(a1-2)Rha(a1-3)Rha(a1-3)[Glc(a1-6)]GlcNAc | GlcNAc(b1-2)Rha(a1-2)Rha(a1-3)Rha2Ac(a1-3)[Glc(a1-6)]GlcNAc | GlcNAc(b1-2)Rha(a1-2)Rha(a1-3)[Glc(a1-4)]Rha(a1-3)GlcNAc | GlcNAc(b1-2)Rha(a1-2)Rha(a1-3)[Glc(b1-4)]Rha(a1-3)GlcNAc | GlcNAc(b1-2)Rha(a1-2)[Glc(a1-3)]Rha(a1-3)Rha(a1-3)GlcNAc | GlcNAc(b1-2)[Glc(a1-3)]Rha(a1-2)Rha(a1-3)Rha(a1-3)GlcNAc | GlcNAc(b1-2)[Glc(a1-3)]Rha(a1-2)Rha(a1-3)Rha2Ac(a1-3)GlcNAc | GlcNAc(b1-2)[Glc(a1-3)]Rha(a1-2)Rha(a1-3)[Glc(a1-4)]Rha(a1-3)GlcNAc | GlcNAc(b1-2)[Glc(a1-3)]Rha(a1-2)[Glc(a1-3)]Rha(a1-3)Rha(a1-3)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-4)][GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-4)][GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Man(a1-3)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-4)]Man(a1-3)[Man(a1-6)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-6)]Man | GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-4)][GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Fuc | GlcNAc(b1-3)FucNAc(a1-4)[GlcNAc(b1-2)]GalA(a1-3)FucNAc(a1-3)GlcNAc | GlcNAc(b1-3)Gal | GlcNAc(b1-3)Gal(b1-3)GalNAc | GlcNAc(b1-3)Gal(b1-3)GlcNAc | GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)]GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)Gal(b1-3)]GalNAc | GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)Gal(b1-6)[Gal(b1-3)]GalNAc | GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | GlcNAc(b1-3)Gal6S(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc | GlcNAc(b1-3)GalNAc | GlcNAc(b1-3)GlcA(a1-4)GlcNAc(a1-3)Rha2Ac(b1-4)GlcNAc | GlcNAc(b1-3)Man | GlcNAc(b1-3)[GalA6CetLys(a1-4)]Gal(a1-3)GlcNAc | GlcNAc(b1-3)[GlcNAc(b1-4)]Gal(b1-4)GlcNAc | GlcNAc(b1-3)[GlcNAc(b1-4)][GlcNAc(b1-6)]GalNAc | GlcNAc(b1-3)[GlcNAc(b1-4)][GlcNAc(b1-6)]GlcNAc | GlcNAc(b1-3)[GlcNAc(b1-6)]Gal | GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-3)[GlcNAc(b1-6)]GalNAc | GlcNAc(b1-3)[GlcNAc(b1-6)]GlcNAc | GlcNAc(b1-3)[GlcNAc(b1-6)]GlcNAc(b1-4)GlcNAc | GlcNAc(b1-3)[Man(b1-4)]Gal(a1-4)Rha(a1-3)GlcNAc | GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)Glc | GlcNAc(b1-3)[Neu5Ac(a2-6)]GalNAc | GlcNAc(b1-4)Fuc | GlcNAc(b1-4)Gal | GlcNAc(b1-4)Gal(b1-4)GlcNAc | GlcNAc(b1-4)GalNAc | GlcNAc(b1-4)GlcA3Ac(b1-2)[Rha3Ac(a1-3)]Man(a1-4)Gal(b1-3)GlcNAc | GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)GlcNAc3Ac | GlcNAc(b1-4)GlcNAc3Prop | GlcNAc(b1-4)GlcNAc6S | GlcNAc(b1-4)Man | GlcNAc(b1-4)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc(b1-4)Mur | GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | GlcNAc(b1-4)[GlcNAc(b1-6)]GalNAc | GlcNAc(b1-6)Gal(b1-4)GlcNAc | GlcNAc(b1-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | GlcNAc(b1-6)GalNAc | GlcNAc(b1-6)Glc(a1-4)[GlcNAc(b1-3)]Gal(b1-3)GalNAc(a1-3)GlcNAc | GlcNAc(b1-6)GlcNAc | GlcNAc(b1-6)Man | GlcNAc(b1-6)Man6P(a1-2)Man(a1-2)Man | GlcNAc(b1-6)Man6P(a1-2)Man(a1-6)Man | GlcNAc(b1-6)Man6P(a1-2)Man(a1-6)Man(a1-6)Man | GlcNAc(b1-6)Man6P(a1-6)Man(a1-6)Man | GlcNAc1P(b1-6)Man(a1-2)Man(a1-2)Man(a1-3)[GlcNAc1P(b1-6)Man(a1-2)Man(a1-6)[Man(a1-2)Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc1P(b1-6)Man(a1-2)Man(a1-3)[GlcNAc1P(b1-6)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc1P(b1-6)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[GlcNAc1P(b1-6)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc1P(b1-6)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc1P(b1-6)Man(a1-2)Man(a1-6)[Man(a1-2)Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc1P(b1-6)Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc2S | GlcNAc2S6S | GlcNAc3Ac(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc3Lac(a1-3)L-QuiNAc(a1-3)GlcNAc(a1-6)GlcNAc3Lac | GlcNAc3Lac(b1-3)Gal(a1-3)GlcNAc6Ac(b1-6)GlcNAc3Lac | GlcNAc3Prop(b1-4)GlcNAc | GlcNAc3S | GlcNAc4Lac(b1-2)Rha(a1-2)Rha(a1-3)[Glc(b1-2)]Rha(a1-3)GlcNAc(b1-3)GlcNAc4Lac | GlcNAc4P(b1-4)Man | GlcNAc4P(b1-6)GlcNAc | GlcNAc4S | GlcNAc6Ac(a1-4)GlcA(a1-3)Fuc(a1-3)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc6Ac | GlcNAc6PEtN(a1-2)Fuc3NBut(b1-6)Glc(a1-4)GlcA(b1-3)GlcNAc6PEtN | GlcNAc6PEtNAc(a1-3)Qui4NAsp2Ac(b1-6)Glc(a1-4)GalA(a1-3)GlcNAc6PEtNAc | GlcNAc6PPenn(a1-3)GlcNAc6PPenn | GlcNAc6S | GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA | GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA | GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA | GlcNAc6S(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA | GlcNAc6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA2S | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA | GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNAc6S(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc6S(b1-2)Man(a1-3)[GlcNAc6S(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | GlcNAc6S(b1-3)Gal(b1-4)GlcNAc | GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S(b1-3)Gal6S | GlcNGc | GlcNS(a1-4)GlcA(b1-4)GlcNS(a1-4)GlcA | GlcNS(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA | GlcNS(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA2S | GlcNS(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA | GlcNS(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNS(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA | GlcNS(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA2S | GlcNS(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA | GlcNS(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNS(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA | GlcNS(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA2S | GlcNS(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA | GlcNS(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNS(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA | GlcNS(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA2S | GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA | GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNS2S | GlcNS2S6S | GlcNS6S | GlcNS6S(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA | GlcNS6S(a1-4)IdoA(a1-4)GlcNAc(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA | GlcNS6S(a1-4)IdoA(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA | GlcNS6S(a1-4)IdoA(a1-4)GlcNS(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA | GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNAc(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS(a1-4)IdoA2S | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA | GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S | GulA | GulA(a1-4)GulA(a1-4)GulA(a1-4)GulA | GulA(a1-4)GulA(a1-4)GulA(a1-4)GulA(a1-4)GulA | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(a1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(a1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(a1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(a1-4)GlcNAc(a1-4)HexA(a1-4)GlcNAc(a1-4)HexA(a1-4)GlcNAc(a1-4)HexA(a1-4)Man | HexA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S(b1-4)GlcA(b1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S(b1-4)IdoA(a1-3)GalNAc4S | HexA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S(b1-4)GlcA(b1-3)GalNAc6S | HexA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc(b1-4)GlcA(b1-3)GlcNAc | HexA2S(a1-4)GlcNS6S | HexA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S | HexA2S(b1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA6S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | HexA6S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA(a1-3)GalNAc | IdoA(a1-3)GalNAc(b1-4)IdoA | IdoA(a1-3)GalNAc4S | IdoA(a1-3)GalNAc4S6S | IdoA(a1-3)GalNAc6S | IdoA(a1-4)GlcN | IdoA(a1-4)GlcN6S | IdoA(a1-4)GlcNAc | IdoA(a1-4)GlcNAc(a1-4)IdoA(a1-4)GlcNAc6S | IdoA(a1-4)GlcNAc(a1-4)IdoA(a1-4)GlcNS | IdoA(a1-4)GlcNAc6S | IdoA(a1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc | IdoA(a1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc6S | IdoA(a1-4)GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc | IdoA(a1-4)GlcNAc6S(a1-4)IdoA(a1-4)GlcNAc6S | IdoA(a1-4)GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc6S | IdoA(a1-4)GlcNS(a1-4)IdoA(a1-4)GlcNAc | IdoA(a1-4)GlcNS(a1-4)IdoA(a1-4)GlcNS | IdoA(a1-4)GlcNS(a1-4)IdoA(a1-4)GlcNS6S | IdoA(a1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA(a1-4)GlcNS3S6S(a1-4)IdoA(a1-4)GlcNS6S | IdoA(a1-4)GlcNS6S | IdoA(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS | IdoA(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | IdoA(a1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | IdoA(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S(b1-4)IdoA(b1-3)GalNAc6S | IdoA(b1-4)Glc6S | IdoA(b1-4)GlcNS6S | IdoA(b1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNS6S | IdoA2S(a1-3)GalNAc4S | IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S(b1-4)IdoA2S(a1-3)GalNAc4S | IdoA2S(a1-3)GalNAc4S6S | IdoA2S(a1-3)GalNAc6S | IdoA2S(a1-4)GlcN | IdoA2S(a1-4)GlcN6S | IdoA2S(a1-4)GlcNAc | IdoA2S(a1-4)GlcNAc(a1-4)GlcA(b1-4)GlcNAc | IdoA2S(a1-4)GlcNAc6S | IdoA2S(a1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc | IdoA2S(a1-4)GlcNAc6S(a1-4)GlcA(b1-4)GlcNAc6S | IdoA2S(a1-4)GlcNAc6S(a1-4)IdoA2S(a1-4)GlcNAc6S | IdoA2S(a1-4)GlcNS | IdoA2S(a1-4)GlcNS(a1-4)GlcA(b1-4)GlcNS | IdoA2S(a1-4)GlcNS6S | IdoA2S(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNAc(a1-4)IdoA2S | IdoA2S(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS | IdoA2S(a1-4)GlcNS6S(a1-4)GlcA(b1-4)GlcNS6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA(a1-4)GlcNAc6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)Man6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)Man6S | IdoA2S(a1-4)GlcNS6S(a1-4)IdoA2S(a1-4)Man6S | IdoA2S(b1-4)Glc6S | IdoA2S(b1-4)Glc6S(a1-4)IdoA2S(a1-4)Glc6S | IdoA2S(b1-4)GlcNS | IdoA2S(b1-4)GlcNS(a1-4)IdoA2S(a1-4)GlcNS | IdoA2S(b1-4)GlcNS6S | IdoA2S(b1-4)GlcNS6S(a1-4)IdoA2S(a1-4)GlcNS6S | IdoA2S4S | Kdn(a2-3)Gal(b1-3)GalNAc | Kdn(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-3)GalNAc(b1-4)[Kdn(a2-3)]Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-3)GlcNAc | Kdn(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-4)GlcNAc | Kdn(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Kdn(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Kdn(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Kdn(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-3)Gal(b1-4)[Rha(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-3)[GalNAc(b1-4)]Gal(b1-3)GlcNAc | Kdn(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Kdn(a2-6)Gal | Kdn(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-6)Gal(b1-4)GlcNAc | Kdn(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn(a2-8)Kdn(a2-3)Gal(b1-4)Glc | Kdn(a2-8)Neu5Ac(a2-3)Gal(b1-4)Glc | Kdn(a2-8)Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Kdn(a2-8)Neu5Gc(a2-3)Gal(b1-4)Glc | Kdn(a2-8)Neu5Gc(a2-3)[Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Kdn(a2-8)Neu5Gc(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Kdn5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Kdn5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Kdn5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Ac9Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Ac9Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Me(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn5Me(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Me(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn5Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn7Me(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn7Me(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn7Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn7Me(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn9Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn9Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn9Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn9Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn9Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdn9Me(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn9Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Kdn9Me(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Kdo | Kdo(a2-4)Kdo | Kdo(a2-8)Kdo | Kdo(a2-8)Kdo(a2-4)Kdo | Kdo(a2-8)Kdo(a2-8)Kdo | Kdo5P | Ko(2-4)Kdo | Ko(a2-4)Kdo | L-Ara | L-Glc | L-Glc(b1-3)FucNAc(a1-3)D-FucNAc(b1-2)L-Glc | L-GulA(b1-4)L-GulA(b1-4)L-GulA(b1-4)L-GulA(b1-4)L-GulA | L-GulNAc3NAmA(a1-4)ManNAc3NAcA(b1-3)D-FucNAc4Ac(a1-4)L-GulNAc3NAmA | L-GulNAcA3NAm(a1-4)ManNAcA3NAc(b1-3)D-FucNAc4Ac(a1-4)L-GulNAcA3NAm | L-PneNAc | L-PneNAc(a1-2)GlcA | L-PneNAc(a1-2)GlcA(b1-3)FucNAc(a1-3)D-FucNAc | L-QuiNAc(a1-3)GlcNAc(a1-4)[L-QuiNAc(a1-3)]GalNAc(a1-4)Gal1P | L-QuiNAc(a1-3)GlcNAc(a1-4)[L-QuiNAc(a1-3)]GalNAc(a1-4)Gal1P(a1-4)L-QuiNAc | L-QuiNAc(a1-3)GlcNAc(a1-6)GlcNAc3Lac(a1-3)L-QuiNAc | LDManHep | LDManHep(a1-2)LDManHep(a1-3)LDManHep(a1-5)Kdo | LDManHep(a1-3)LDManHep | LDManHep(a1-3)LDManHep(a1-5)Kdo | LDManHep(a1-3)LDManHep(a1-5)[Ara4N(b1-8)]Kdo | LDManHep(a1-5)Kdo | LDManHep(a1-5)Kdo4P | LDManHep(a1-7)LDManHep(a1-3)LDManHep | LDManHep(a1-7)LDManHep(a1-3)LDManHep(a1-5)Kdo | Leg5Ac7Ala(a2-4)GlcA(b1-3)GlcNAc(b1-8)Leg5Ac7Ala | Man | Man(a1-2)Fuc(a1-2)GlcA4Ac(b1-3)GalNAc(a1-3)Man | Man(a1-2)Man | Man(a1-2)Man(a1-2)Man | Man(a1-2)Man(a1-2)Man(a1-2)Man | Man(a1-2)Man(a1-2)Man(a1-3)Man | Man(a1-2)Man(a1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[GlcNAc1P(b1-6)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man6P(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-3)[Man6P(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man6P(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-2)Man(a1-3)[Man6P(a1-6)]Man | Man(a1-2)Man(a1-2)Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-2)Man(b1-3)GalNAc(a1-4)Man | Man(a1-2)Man(a1-2)[Gal(b1-4)]Man | Man(a1-2)Man(a1-2)[Man(a1-2)Man(a1-6)]Man | Man(a1-2)Man(a1-2)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)Man | Man(a1-2)Man(a1-3)Man(a1-6)Man | Man(a1-2)Man(a1-3)Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-3)Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-3)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)Man | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-3)[Man(a1-6)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man | Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-3)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-3)[Man6P(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man | Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)Man | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[GlcNAc1P(b1-6)Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(a1-6)[Man6P(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-3)[Man4P(a1-6)]Man | Man(a1-2)Man(a1-3)[Man6P(a1-6)Man(a1-6)]Man | Man(a1-2)Man(a1-3)[Man6P(a1-6)]Man | Man(a1-2)Man(a1-3)[Man6P(a1-6)]Man(a1-6)[Man6P(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-6)Man | Man(a1-2)Man(a1-6)Man(a1-6)Man | Man(a1-2)Man(a1-6)Man(a1-6)Man(b1-4)GlcNAc | Man(a1-2)Man(a1-6)Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-6)Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-6)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-2)Man(a1-6)[GalNAc(b1-4)]Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)Man(b1-4)GlcNAc | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-3)]Man | Man(a1-2)Man(a1-6)[Man(a1-3)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-2)[Glc(a1-3)]Man(a1-2)Man(a1-2)Man(b1-3)GlcNAc(a1-6)[Rha(a1-3)]Man | Man(a1-2)[Glc(a1-4)]Man(a1-2)[Glc(a1-3)]Man(b1-3)GlcNAc(a1-6)Man | Man(a1-2)[Glc(a1-4)]Man(b1-3)GlcNAc(a1-6)Man(a1-2)Man | Man(a1-2)[Man(a1-3)]Man(a1-6)Man(b1-4)GlcNAc | Man(a1-2)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-3)6dTal(a1-3)GalNAc(b1-4)[6dTal(a1-3)]Man | Man(a1-3)Fuc(a1-3)GalNAc(a1-4)[Gal(b1-3)]GalNAc(b1-2)[Glc(a1-3)]Man | Man(a1-3)Fuc(a1-3)GlcNAc(b1-4)[Rha(a1-2)Fuc(a1-3)]Man | Man(a1-3)L-6dTal(a1-3)GalNAc(b1-4)[6dTal(a1-3)]Man | Man(a1-3)Man | Man(a1-3)Man(a1-6)Man | Man(a1-3)Man(a1-6)Man(b1-4)GlcNAc | Man(a1-3)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(a1-6)[Man(a1-3)]Man | Man(a1-3)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-3)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)Man(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)GlcNAc | Man(a1-3)Man(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)Man(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Man(a1-3)Man(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Man(a1-3)[Man(a1-4)][Man(a1-6)]Man | Man(a1-3)[Man(a1-6)]Man | Man(a1-3)[Man(a1-6)]Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)Man | Man(a1-3)[Man(a1-6)]Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Man(a1-3)[Man6P(a1-6)]Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(a1-4)Man | Man(a1-4)Rha(a1-3)GlcNAc(a1-3)Qui4NBut(b1-4)[Gal(a1-3)]Man | Man(a1-4)[Man(a1-6)]Man | Man(a1-5)Araf(a1-3)[Man(a1-5)Araf(a1-5)]Araf(a1-5)Araf | Man(a1-6)Man | Man(a1-6)Man(a1-4)[Man(a1-6)Man(a1-6)]Man | Man(a1-6)Man(a1-6)Man | Man(a1-6)Man(a1-6)Man(b1-4)GlcNAc | Man(a1-6)Man(a1-6)[Man(a1-3)]Man | Man(a1-6)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc | Man(a1-6)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Gal(b1-4)Fuc(a1-6)][Fuc(a1-3)]GlcNAc | Man(a1-6)Man(a1-6)[Man6P(a1-3)]Man | Man(a1-6)Man(b1-4)GlcNAc | Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(b1-3)Gal(a1-4)Rha(a1-3)GlcNAc(a1-2)[RhaNAc3NFo(b1-3)]Man | Man(b1-3)Glc(b1-3)GlcNAc(b1-2)[Fuc3NFo(a1-3)]Man | Man(b1-4)Glc(a1-3)QuiNAc(a1-3)GlcNAc(a1-2)Man | Man(b1-4)GlcA(b1-3)L-QuiNAc(a1-3)GlcNAc(a1-2)Man | Man(b1-4)GlcNAc | Man(b1-4)GlcNAc(b1-4)GlcNAc | Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Man(b1-4)Man(a1-3)GalNAc(b1-4)[Rha2Ac3Lac(a1-3)]Man | Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man(b1-4)Man | Man6P | Man6P(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man6P(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man6P(a1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man6P(a1-2)Man(a1-3)[Man(a1-3)[Man6P(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Man6P(a1-3)Man(a1-3)Man(a1-3)Man(a1-2)Man | Man6P(a1-6)Man(a1-6)[Man(a1-3)]Man | Man6PEtN(a1-2)Man(a1-6)[GalNAc(b1-4)]Man | Man6PEtN(a1-2)Man(a1-6)[GalNAc(b1-4)]Man2PEtN | ManA | ManA(b1-4)ManA(b1-4)ManA(b1-4)ManA | ManA(b1-4)ManA(b1-4)ManA(b1-4)ManA(b1-4)ManA | ManNAc | ManNAc(b1-3)FucNAc(a1-3)GalNAc(a1-4)Gal | ManNAc(b1-3)FucNAc(a1-3)GalNAc(a1-4)Gal3Pyr | ManNAc(b1-4)Glc(b1-4)Glc | ManNAc3NAcA(b1-2)Rha(a1-3)Rha(b1-4)GlcNAc(a1-4)ManNAc3NAcA | ManNAc3NAmA(b1-4)L-GulNAc3NAcA(a1-3)D-FucNAc(b1-4)ManNAc3NAmA | ManNAc3NAmA(b1-4)L-GulNAc3NAcA(a1-3)D-FucNAc4N(b1-4)ManNAc3NAmA | ManNAc3NAmA(b1-4)ManNAc3NAcA(b1-3)D-FucNAc(a1-4)ManNAc3NAmA | ManNAcA3NAc(b1-2)Rha(a1-3)Rha(b1-4)GlcNAc(a1-4)ManNAcA3NAc | ManNAcA3NAm(b1-4)L-GulNAcA3NAc(a1-3)D-FucNAc(b1-4)ManNAcA3NAm | ManNAcA3NAm(b1-4)L-GulNAcA3NAc(a1-3)D-FucNAc4N(b1-4)ManNAcA3NAm | ManNAcA3NAm(b1-4)ManNAcA3NAc(b1-3)D-FucNAc(a1-4)ManNAcA3NAm | MurNAc(a1-4)GlcNAc | MurNAc(b1-4)GlcNAc | Neu(a2-3)Gal(b1-4)Glc | Neu(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu(a2-3)Gal(b1-4)GlcNAc6S(b1-3)Gal(b1-4)Glc | Neu(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc6S(b1-3)Gal(b1-4)Glc | Neu(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcN(b1-3)Gal(b1-4)Glc | Neu(a2-6)Gal(b1-4)Glc | Neu4Ac5Ac(a2-3)Gal(b1-4)Glc | Neu4Ac5Ac(a2-3)Gal(b1-4)GlcNAc | Neu4Ac5Ac(a2-6)Gal(b1-4)GlcNAc | Neu4Ac5Ac7Ac9Ac(a2-3)Gal(b1-4)GlcNAc | Neu4Ac5Ac7Ac9Ac(a2-6)Gal(b1-4)GlcNAc | Neu4Ac5Ac9Ac(a2-3)Gal(b1-4)GlcNAc | Neu4Ac5Ac9Ac(a2-6)Gal(b1-4)GlcNAc | Neu4Me5Ac(a2-3)Gal(b1-4)Glc | Neu4S5Ac(a2-3)Gal(b1-4)GlcNAc | Neu4S5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu4S5Ac(a2-3)Gal6S(b1-4)GlcNAc | Neu4S5Ac(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcNAc | Neu5A(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5A(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5A(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac | Neu5Ac(a2-2)Glc | Neu5Ac(a2-2)[Neu5Ac(a2-3)]Gal | Neu5Ac(a2-3)4dGal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)6dGal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)FucNAm(a1-3)GlcNAc6Ac(b1-7)Neu5Ac | Neu5Ac(a2-3)Gal | Neu5Ac(a2-3)Gal(a1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(a1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)6dGalNAc | Neu5Ac(a2-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)Gal3S(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc6S | Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)Gal3S(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-4)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-3)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc(b1-6)GalNAc | Neu5Ac(a2-3)Gal(b1-3)GlcNAc6S | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-3)]GlcN(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-3)Gal(b1-4)]GlcNAc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal3S(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-6)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)]Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)GalNAc(a1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-3)Gal6S(b1-3)GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-3)Gal6S(b1-4)GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)ManNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[3dFuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[4dFuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[6dTal(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-4)GalNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Neu5Ac(a2-3)Gal(b1-3)]GalNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc6S(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc6S(b1-3)Gal6S(b1-3)GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-4)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-4)]GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)Gal(b1-4)[Fuc(b1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc2Me(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[Fuc2S(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Fuc2S(a1-3)]GlcNAc6S | Neu5Ac(a2-3)Gal(b1-4)[Fuc3S(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal(b1-4)[Qui(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal(b1-4)[Rha(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac(a2-3)Gal3S | Neu5Ac(a2-3)Gal6S | Neu5Ac(a2-3)Gal6S(b1-3)GalNAc | Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc | Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-3)Gal6S(b1-4)GlcNAc6S | Neu5Ac(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcNAc6S(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)GalNAc | Neu5Ac(a2-3)GalNAc(b1-4)GlcNAc | Neu5Ac(a2-3)GalNBz(b1-4)GlcNAc | Neu5Ac(a2-3)Glc | Neu5Ac(a2-3)[Gal(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-3)Gal | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-3)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-6)GalNAc | Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-3)]GalNAc | Neu5Ac(a2-3)[GalNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-3)[GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-3)[Neu5Ac(a2-6)]Gal(b1-3)GalNAc | Neu5Ac(a2-3)[Neu5Ac(a2-6)]GalNAc | Neu5Ac(a2-6)Gal | Neu5Ac(a2-6)Gal(a1-3)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-3)GlcNAc6S | Neu5Ac(a2-6)Gal(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-3)[GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[GlcNAc(b1-4)][Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)]Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)]Man(a1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-2)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Fuc(a1-3)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-2)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Ac(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)]Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-4)Man(a1-6)][GlcNAc(b1-4)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-6)Gal(b1-3)]GalNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc6S | Neu5Ac(a2-6)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc6S(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal(b1-6)GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-6)Gal3S(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal3S(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-6)Gal3S(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)Gal6S | Neu5Ac(a2-6)Gal6S(b1-4)Glc | Neu5Ac(a2-6)Gal6S(b1-4)Glc4S | Neu5Ac(a2-6)GalNAc | Neu5Ac(a2-6)GalNAc(b1-4)GlcNAc | Neu5Ac(a2-6)GalNAc(b1-4)GlcNAc6S | Neu5Ac(a2-6)Glc | Neu5Ac(a2-6)Glc6S | Neu5Ac(a2-6)GlcNAc | Neu5Ac(a2-6)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)GlcNAc(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-6)GlcNAc6S | Neu5Ac(a2-8)Kdn(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)Gal3S(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)Gal3S(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)[Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal(b1-3)GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-3)[GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-6)Gal | Neu5Ac(a2-8)Neu5Ac(a2-6)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-8)Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-8)Neu5Ac(a2-6)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal(b1-3)GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Gal3S(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Neu5Ac(a2-3)Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Neu5Ac(a2-3)Gal(b1-3)GalNAc6S(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-6)Gal | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac(a2-8)Neu5Ac | Neu5Ac(a2-8)Neu5Gc(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc | Neu5Ac(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-8)Neu5Gc(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Ac(a2-8)Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac(a2-8)Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac(a2-9)Neu5Ac | Neu5Ac(a2-9)Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac | Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac | Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac | Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac(a2-9)Neu5Ac | Neu5Ac(b1-6)GalNAc | Neu5Ac(b2-3)Gal(b1-4)Glc | Neu5Ac(b2-3)Gal(b1-4)GlcNAc | Neu5Ac(b2-6)Gal | Neu5Ac(b2-6)Gal(b1-3)GalNAc | Neu5Ac(b2-6)Gal(b1-4)Glc | Neu5Ac(b2-6)Gal(b1-4)GlcNAc | Neu5Ac(b2-6)GalNAc | Neu5Ac4S(a2-3)Gal6S(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac7Ac(a2-6)Gal(b1-4)GlcNAc | Neu5Ac7Ac9Ac(a2-3)Gal(b1-4)GlcNAc | Neu5Ac7Ac9Ac(a2-6)Gal(b1-4)GlcNAc | Neu5Ac8Me(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac8Me(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac8Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac8Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac8Me(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Ac | Neu5Ac9Ac(a2-3)Gal(b1-3)GlcNAc | Neu5Ac9Ac(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Ac(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Neu5Ac9Ac(a2-3)Gal(b1-4)GlcNAc | Neu5Ac9Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Ac9Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Ac9Ac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Ac(a2-6)Gal(b1-4)GlcNAc | Neu5Ac9Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac9Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Ac(a2-8)Neu5Ac | Neu5Ac9Lac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac9Lac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Lac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Lac(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Lac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Ac9Lac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Lac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Me(a2-3)Gal(b1-4)Glc | Neu5Ac9Me(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Me(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9Me(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Ac9Me(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Ac9S(a2-3)Gal(b1-4)GlcNAc | Neu5Gc | Neu5Gc(a2-3)Gal | Neu5Gc(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-3)GalNAc(b1-3)Gal(a1-4)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-3)GlcNAc | Neu5Gc(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-3)GlcNAc6S | Neu5Gc(a2-3)Gal(b1-3)[Fuc(a1-4)]GlcNAc | Neu5Gc(a2-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)GlcNAc6S | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcA(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcA3S(b1-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)[Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-6)[GlcNAc(b1-2)]Man | Neu5Gc(a2-3)Gal(b1-4)[Fuc(a1-4)]GlcNAc | Neu5Gc(a2-3)[GalNAc(b1-4)]Gal | Neu5Gc(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-3)GlcNAc | Neu5Gc(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)[GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-6)]Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Gc(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-3)[Neu5Gc(a2-6)]Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-4)GlcNAc(b1-3)]Gal(b1-4)Glc | Neu5Gc(a2-6)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-6)GalNAc | Neu5Gc(a2-8)Neu5Ac(a2-3)Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Ac(a2-3)[Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Ac(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-8)Neu5Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Gc(a2-3)Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Gc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Gc(a2-3)[Gal(b1-3)GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Gc(a2-3)[GalNAc(b1-4)]Gal(b1-4)Glc | Neu5Gc(a2-8)Neu5Gc(a2-6)Gal(b1-4)GlcNAc | Neu5Gc(a2-8)Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc(a2-8)Neu5Gc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc(b2-6)Gal(b1-4)GlcNAc | Neu5Gc(b2-6)GalNAc | Neu5Gc9Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc9Ac(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc9Ac(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5Gc9Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5Gc9Ac(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5Gc9Ac(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOAc(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5GcOAc(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOAc(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOAc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5GcOAc(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5GcOAc(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOMe(a2-3)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOMe(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5GcOMe(a2-3)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5GcOMe(a2-3)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOMe(a2-6)Gal(b1-3)GlcNAc(b1-3)Gal(b1-4)Glc | Neu5GcOMe(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Neu5GcOMe(a2-6)Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Neu5GcOMe(a2-6)Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc | Pse4Ac5Ac7But(b2-4)D-FucNAc(a1-3)QuiNAc(b1-3)Pse4Ac5Ac7But | Qui3N(a1-3)Rha(a1-4)Gal(b1-3)[Glc(b1-4)]GalN(a1-4)Qui3N | Qui3NAc(a1-3)Rha(a1-4)Gal(b1-3)GalNAc(a1-4)Qui3NAc | Qui3NAc(a1-3)Rha(a1-4)Gal(b1-3)[Glc(b1-4)]GalNAc(a1-4)Qui3NAc | Qui3NAc(a1-3)Rha(a1-6)GlcNAc(a1-4)GlcA(b1-3)[Glc(b1-4)]GalNAc(a1-4)Qui3NAc | Qui3NFo(b1-3)Gal(a1-3)GlcA(b1-3)GalNAc(b1-4)Qui3NFo | Qui4NAsp2Ac(b1-6)Glc(a1-4)GalA(a1-3)[GlcNAc6PEtNAc(a1-3)]Qui4NAsp2Ac | Qui4NBut(a1-4)GalNAc(b1-4)Rha(a1-3)GlcNAc6Ac(b1-2)Qui4NBut | Qui4NButAla(b1-6)GlcNAc(a1-3)L-QuiNAc(a1-3)GlcNAc6Ac(a1-3)Qui4NButAla | Qui4NButAla(b1-6)GlcNAc(a1-3)QuiNAc(a1-3)GlcNAc(a1-3)Qui4NButAla | Qui4NSer2Ac(b1-3)Ribf(b1-4)GalNAc(a1-3)GlcNAc(a1-3)Qui4NSer2Ac | Qui4NSer2Ac(b1-4)Ribf(b1-4)GalNAc(a1-3)GlcNAc(a1-3)Qui4NSer2Ac | Rha | Rha(a1-2)Rha(a1-2)Gal(a1-3)GlcNAc(b1-3)[ManNAc(b1-2)]Rha | Rha(a1-2)Rha(a1-2)Gal6Ac(a1-3)GlcNAc(b1-3)[Glc(b1-3)GlcNAc4Lac(b1-2)]Rha | Rha(a1-2)Rha(a1-2)Rha | Rha(a1-2)Rha(a1-2)Rha(a1-2)Glc(a1-3)GlcNAc6Ac(a1-2)Rha | Rha(a1-2)Rha(a1-2)Rha(a1-3)[Glc(a1-2)]Rha(a1-3)[Glc(a1-2)]Rha | Rha(a1-2)Rha(a1-2)Rha(a1-4)GalA(a1-3)GlcNAc(a1-2)Rha | Rha(a1-2)Rha(a1-3)Gal(a1-3)D-FucNAc(a1-3)Rha(b1-4)[Glc(a1-3)]Rha | Rha(a1-2)Rha(a1-3)Rha(a1-2)Rha(a1-3)GlcNAc(b1-3)[GalNAcA6N(a1-2)]Rha | Rha(a1-2)Rha(a1-3)Rha(a1-3)GlcNAc(b1-2)Rha | Rha(a1-2)Rha(a1-3)Rha(a1-3)[Glc(a1-4)]GlcNAc(b1-2)[Glc(a1-3)]Rha | Rha(a1-2)Rha(a1-3)Rha2Ac(a1-3)GlcNAc(b1-2)Rha | Rha(a1-2)Rha(a1-3)Rha2Ac(a1-3)[Glc(a1-6)]GlcNAc(b1-2)Rha | Rha(a1-2)Rha(a1-4)Glc(a1-3)GalNAc(b1-3)[GlcNAc(b1-2)]Rha | Rha(a1-2)Rha3PEtN(a1-3)Rha(a1-3)GlcNAc(b1-2)[Glc(a1-3)]Rha | Rha(a1-3)FucNAc(a1-3)FucNAc(a1-3)D-FucNAc(a1-2)Rha | Rha(a1-3)FucNAc(a1-3)FucNAc(a1-3)QuiNAc(a1-2)Rha | Rha(a1-3)Glc | Rha(a1-3)Glc(b1-4)Glc | Rha(a1-3)Glc(b1-4)[Rha(a1-3)]Glc(a1-2)Glc | Rha(a1-3)Rha(a1-2)Gal(a1-3)GlcNAc(a1-3)Rha | Rha(a1-3)Rha(a1-2)Glc(a1-3)GlcNAc(a1-3)Rha | Rha(a1-3)Rha(a1-3)Rha(a1-3)GalNAc(a1-2)[ManNAc(a1-3)]Rha | Rha(a1-3)Rha(a1-4)GalNAcA(a1-3)QuiNAc(b1-3)Rha(a1-3)[Glc(a1-6)]Glc(b1-3)[Glc(a1-4)]GalN2Ala(a1-3)LDManHep6P7Cm(a1-3)LDManHep2P4P(a1-5)Kdo | Rha(a1-3)Rha(a1-4)GalNAcA3Ac(a1-3)QuiNAc(b1-2)Rha | Rha(a1-3)[Glc(a1-4)]GlcNAc(b1-2)Rha(a1-2)Rha(a1-3)Rha | Rha(a1-4)GalNAcA3Ac(a1-4)GalNFoA(a1-3)QuiNAc(a1-2)Rha | Rha(a1-4)Glc(a1-2)Rha(a1-3)GlcNAc(b1-3)[Rha2Ac3Ac4Ac(a1-4)GalA(a1-2)]Rha | Rha(a1-4)L-GalNAcA(a1-3)QuiNAc(a1-3)Rha | Rha(a1-6)Glc | Rha(b1-4)Rha(a1-2)Rha(a1-2)GalA(a1-3)GalNAc(a1-3)Rha | Rha(b1-4)Rha(a1-4)GlcA(b1-3)GlcNAc(b1-3)[Qui2Ac3NAc4Ac(a1-2)]Rha | Rha3Ac(a1-2)Rha(a1-3)Rha(a1-3)GlcNAc6Ac(b1-2)Rha | Rha3Ac(a1-2)Rha(a1-3)Rha(a1-3)GlcNAc6Ac(b1-2)Rha3Ac | Rha3Ac(a1-2)Rha(a1-4)GalA(b1-3)GalNAc(b1-2)Rha3Ac | Rha3Ac(a1-6)GlcNAc(a1-4)L-GalNAcA(a1-3)QuiNAc4NHb(b1-2)Rha3Ac | Rha3Ac(a1-6)GlcNAc(a1-4)L-GalNAcA(a1-3)QuiNAc4NSHb(b1-2)Rha3Ac | Rha3Ac(a1-6)GlcNAc(a1-4)L-GalNAcA3Ac(a1-3)QuiNAc4NBut(b1-2)Rha3Ac | Rha3Ac4Ac(a1-2)Rha(a1-4)GalA(b1-3)GalNAc(b1-2)Rha3Ac4Ac | Rha3PEtN(a1-2)Rha(a1-3)Rha(a1-3)[Glc(a1-6)]GlcNAc(b1-2)Rha3PEtN | Rha3PEtN(a1-2)Rha3PEtN(a1-3)Rha(a1-3)GlcNAc(b1-2)Rha3PEtN | Rha3PEtN(a1-2)Rha3PEtN(a1-3)Rha(a1-3)GlcNAc6Ac(b1-2)Rha3PEtN | Rha4NAc(a1-3)Fuc(a1-4)Glc(b1-3)GalNAc(a1-2)Rha4NAc | Rib | Rib1Ac5P-ol(5-6)Gal(b1-4)[GlcNAc(b1-2)]Glc(b1-3)GlcNAc(b1-3)Rib1Ac5P-ol(5-6)Gal | Rib5P-ol(5-3)Gal(b1-3)GlcNAc(a1-3)Glc6PEtN(b1-3)GlcNAc(b1-2)Rib5P-ol(5-3)Gal | Rib5P-ol(5-6)Gal(a1-3)FucNAm(a1-3)GlcN(b1-2)Rib5P-ol(5-6)Gal | Rib5P-ol(5-6)Gal(a1-3)FucNAm(a1-3)GlcNAc(b1-3)Rib5P-ol(5-6)Gal | Rib5P-ol(5-6)Gal(a1-3)FucNAm(a1-3)[GlcNAc(b1-4)]GlcNAc(b1-2)Rib5P-ol(5-6)Gal | Ribf(b1-3)GalNAc(a1-2)Ribf | Ribf(b1-4)Gal(b1-4)GlcNAc(a1-4)Gal(a1-3)GlcNAc(a1-2)Ribf | Ribf(b1-4)Gal(b1-4)GlcNAc(a1-4)Gal(b1-3)GlcNAc(a1-2)Ribf | Ribf(b1-4)GalA(a1-3)GlcNAc(a1-2)[Galf(b1-3)]Rha(a1-2)Rha(a1-2)Ribf(b1-4)GalA(a1-2)Ribf | Xyl | Xyl(a1-3)Glc | Xyl(a1-3)Xyl(a1-3)Glc | Xyl(a1-6)Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)[Xyl(a1-6)]Glc(b1-4)Glc | Xyl(b1-2)Man | Xyl(b1-2)Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Xyl(b1-2)[Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-3)][Fuc(a1-6)]GlcNAc | Xyl(b1-2)[Man(a1-3)][Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Xyl(b1-2)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc | Xyl(b1-2)[Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc | Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl(b1-4)Xyl | target | protein |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.293462 | -1.316793 | nan | -0.860744 | -1.211838 | -0.335253 | -1.127330 | nan | nan | -0.070201 | nan | -0.262080 | -0.555655 | -1.313299 | nan | -0.134637 | nan | nan | nan | nan | nan | nan | 0.317005 | 0.821230 | nan | nan | nan | -1.076498 | nan | -0.068528 | 1.830645 | nan | -0.670428 | nan | 0.691076 | nan | 0.276549 | nan | nan | nan | nan | nan | -1.170653 | 0.295167 | nan | nan | nan | 0.401201 | -1.117411 | 1.983681 | -0.524707 | -0.792918 | 0.556915 | nan | 0.281728 | nan | nan | nan | nan | nan | 0.858230 | 0.750740 | nan | -0.326871 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.490199 | nan | nan | nan | -0.019968 | 0.206706 | nan | nan | nan | nan | nan | nan | -0.321137 | nan | nan | nan | -0.315444 | nan | nan | nan | nan | -0.061100 | -1.022834 | 0.778259 | nan | -1.122821 | nan | nan | nan | nan | nan | nan | -0.866634 | 0.898772 | nan | nan | nan | 0.753109 | nan | nan | nan | nan | nan | nan | -0.570608 | nan | nan | nan | nan | -0.370622 | -0.214434 | -0.083591 | -0.202393 | -0.234208 | nan | nan | 0.228359 | nan | -0.474023 | nan | -0.130498 | nan | -0.599990 | -0.336374 | 1.158122 | nan | nan | -1.315247 | -0.942153 | nan | 1.501823 | -0.334507 | -0.251028 | 1.006794 | -0.859050 | nan | nan | nan | nan | -1.982244 | nan | -0.676445 | 0.471350 | 0.036644 | 0.468515 | -1.810952 | nan | 2.315709 | nan | nan | nan | 0.199587 | nan | nan | nan | -0.505418 | nan | 0.728243 | nan | 0.026061 | 0.321230 | -1.398725 | nan | nan | nan | 0.334855 | -1.018204 | nan | nan | -0.620614 | 0.033777 | 1.862801 | -0.315860 | 0.550124 | nan | nan | nan | -0.013552 | nan | -0.509655 | nan | -0.417772 | -0.156735 | -0.549880 | nan | nan | -0.161960 | -0.731572 | nan | -0.855801 | -0.491026 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.555533 | nan | nan | nan | nan | nan | nan | nan | nan | 0.194983 | 0.512153 | -0.996060 | nan | 0.948385 | 0.618666 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.454668 | nan | nan | nan | nan | nan | nan | nan | -0.321611 | nan | nan | nan | -1.673173 | nan | nan | nan | nan | -0.605793 | nan | nan | nan | nan | nan | nan | nan | nan | -0.062013 | nan | nan | nan | nan | nan | 0.659781 | -0.287028 | 0.417449 | -0.293227 | nan | nan | 0.585646 | nan | nan | nan | 3.189814 | nan | nan | nan | nan | nan | -0.396293 | -0.530146 | nan | nan | nan | 1.141278 | nan | nan | nan | nan | -0.301479 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.206983 | nan | nan | -0.661388 | nan | -1.726143 | -0.809143 | 0.359409 | -1.036932 | 0.369392 | 0.387420 | nan | -0.806718 | -0.344472 | -1.688612 | nan | -0.266892 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.345571 | nan | nan | nan | nan | nan | -1.348697 | nan | nan | nan | 0.730832 | -1.557567 | nan | nan | nan | -0.635470 | nan | nan | nan | nan | nan | -0.656538 | nan | nan | nan | nan | nan | -0.355969 | nan | nan | -0.551805 | 0.196506 | 0.170764 | nan | -1.291785 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 2.593879 | nan | 1.790022 | nan | nan | nan | nan | nan | nan | 0.734081 | nan | nan | nan | nan | nan | nan | -0.511112 | nan | nan | nan | nan | 2.920829 | nan | nan | -0.084440 | nan | -0.472743 | -0.542256 | nan | -1.256444 | nan | -0.155544 | nan | nan | nan | nan | 0.177562 | nan | nan | nan | nan | 0.291092 | nan | nan | nan | nan | nan | nan | nan | -0.678792 | nan | -0.809929 | nan | nan | nan | 0.826958 | -0.520985 | -1.125411 | -0.545094 | 1.177174 | 0.473622 | nan | nan | nan | nan | nan | -0.096637 | nan | 1.221971 | -1.017643 | 0.182799 | -0.348565 | nan | nan | 0.205954 | nan | nan | nan | nan | -0.934454 | nan | nan | -0.408113 | nan | nan | nan | -0.224584 | nan | 0.662677 | 0.549441 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.002986 | -0.262705 | -0.366825 | nan | 3.147103 | nan | nan | nan | nan | nan | nan | nan | -0.073753 | 0.506367 | nan | 1.229815 | nan | -0.763115 | nan | nan | nan | 0.013598 | nan | nan | nan | nan | nan | -0.100337 | nan | nan | nan | 2.391141 | nan | -0.969170 | nan | nan | nan | nan | nan | nan | nan | nan | -0.310658 | nan | -0.444067 | nan | nan | -1.420922 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.418361 | 1.068793 | -0.134195 | nan | nan | -0.333582 | 0.513704 | nan | 0.443801 | nan | 0.361051 | 0.990320 | nan | nan | 0.345317 | nan | 0.685388 | -0.637602 | -0.531655 | -0.903252 | nan | -0.699069 | 0.291485 | -1.058475 | 0.960072 | nan | nan | nan | nan | 0.205642 | nan | nan | nan | nan | 1.102914 | 0.194423 | nan | nan | nan | nan | nan | nan | -0.255641 | nan | nan | nan | nan | nan | nan | nan | 3.527070 | nan | -0.088100 | -1.013221 | nan | -0.822409 | -0.429073 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.484419 | -0.822507 | -0.778704 | 0.031066 | 0.102814 | -0.013922 | nan | -0.104233 | 0.235792 | nan | nan | nan | -0.692381 | nan | nan | nan | nan | -0.371819 | 0.791884 | -0.238029 | nan | nan | nan | nan | nan | nan | -1.341460 | 0.567337 | 0.311051 | nan | nan | nan | -0.739965 | 1.503106 | -0.777380 | 0.224688 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.204058 | nan | nan | 0.880339 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.169474 | nan | nan | nan | nan | nan | nan | nan | -0.175544 | nan | nan | nan | nan | nan | nan | -0.186680 | -1.767021 | nan | nan | nan | nan | 0.074453 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.866123 | nan | 0.861154 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.275722 | nan | nan | 0.760378 | nan | -0.122172 | nan | nan | nan | nan | nan | 1.528204 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.059112 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.266091 | nan | nan | nan | nan | nan | nan | 1.919748 | nan | nan | -1.299860 | nan | nan | nan | nan | 1.820241 | 0.909694 | -0.394599 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.189469 | 2.275547 | 1.879986 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.034795 | nan | 1.943014 | nan | nan | nan | nan | nan | 0.132822 | nan | nan | nan | -1.320108 | 0.017783 | nan | nan | nan | -0.536308 | 2.232935 | 0.993916 | nan | nan | nan | nan | -0.398402 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.613325 | nan | nan | nan | nan | nan | 0.657140 | nan | nan | nan | 0.083962 | 0.070875 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.639347 | nan | nan | 0.963136 | 0.340265 | -0.435801 | nan | nan | 1.314698 | nan | -0.707144 | nan | nan | nan | nan | nan | 0.010442 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.409104 | 1.312975 | nan | -0.647214 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.378275 | nan | nan | nan | 0.091274 | nan | nan | -1.098850 | nan | nan | nan | nan | nan | nan | nan | nan | -0.087915 | nan | -1.070024 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.161683 | nan | nan | nan | nan | nan | nan | -0.544354 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.583946 | nan | -0.205191 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.264347 | -2.010649 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.293906 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.845928 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.252531 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.370796 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.729693 | nan | nan | nan | nan | nan | nan | -1.828778 | nan | nan | nan | nan | nan | nan | nan | 1.019621 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.163787 | nan | nan | nan | nan | nan | nan | nan | -0.118932 | nan | -0.209555 | 0.133655 | -0.136596 | 0.335849 | -0.673942 | 0.090147 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.037059 | nan | nan | 0.736757 | nan | nan | nan | nan | nan | nan | 2.788943 | nan | nan | 0.725185 | 0.424761 | -0.145538 | nan | nan | nan | nan | 0.370871 | -1.507851 | -0.447876 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.841155 | nan | -0.902281 | nan | nan | nan | nan | nan | -1.070313 | -0.549892 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.387773 | nan | -0.060701 | 0.173437 | nan | -1.081284 | -0.603117 | -0.266577 | -0.050777 | 0.402848 | nan | -0.523505 | nan | nan | nan | 0.029309 | -0.717797 | -0.522505 | nan | nan | nan | 0.114510 | -0.156128 | -0.597342 | nan | 1.445367 | -0.014696 | -0.450986 | nan | 0.580938 | nan | nan | nan | -0.866310 | nan | nan | -0.152070 | 0.586291 | -0.887316 | -0.090753 | nan | nan | nan | nan | nan | nan | -0.740103 | nan | 0.386357 | nan | nan | nan | 0.432027 | nan | nan | 0.229075 | nan | -0.104972 | nan | nan | nan | nan | nan | nan | nan | 0.158284 | nan | nan | -0.600516 | -1.672901 | nan | 1.439656 | -0.051135 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.968090 | -0.938656 | nan | -0.417570 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.390362 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.561521 | nan | nan | nan | 0.509708 | nan | 0.121435 | 0.399651 | nan | nan | -0.466956 | nan | nan | nan | nan | nan | nan | nan | -0.602359 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.592597 | nan | nan | nan | nan | 0.042089 | -0.832484 | nan | nan | nan | nan | nan | 1.375992 | -1.149122 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.189469 | nan | nan | nan | nan | -1.230641 | nan | nan | nan | nan | nan | nan | 0.034991 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.280536 | nan | nan | nan | nan | nan | nan | nan | nan | 0.599822 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 3.240981 | nan | nan | nan | nan | nan | nan | nan | nan | -0.512395 | -0.006985 | nan | nan | nan | -0.368857 | nan | nan | 0.616198 | nan | nan | nan | nan | nan | -1.365917 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.015448 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.836073 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.938211 | nan | nan | nan | -0.869553 | nan | nan | 1.554984 | nan | nan | nan | 0.854212 | 0.392172 | 0.743144 | nan | nan | nan | nan | 0.415767 | -0.188451 | nan | 0.679053 | -0.322970 | nan | nan | nan | nan | -0.898535 | nan | -0.568447 | nan | 0.525800 | nan | nan | -0.389333 | -0.944222 | -0.021667 | nan | nan | -1.095452 | 1.406847 | nan | 1.647619 | nan | -0.435847 | 0.718515 | 0.082123 | nan | nan | nan | nan | nan | nan | nan | -1.215352 | nan | nan | nan | 1.217300 | -0.435636 | -0.663364 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.201278 | -0.509222 | -0.429130 | 0.740803 | nan | -0.364258 | nan | nan | nan | -0.106666 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.167387 | 0.340328 | nan | nan | nan | nan | nan | nan | -1.250190 | nan | -0.878616 | nan | nan | nan | nan | 0.077389 | nan | nan | 2.517735 | nan | nan | nan | nan | nan | -0.019361 | nan | nan | nan | nan | nan | nan | nan | 1.491176 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.303617 | -0.672405 | 0.317837 | 0.332005 | nan | nan | nan | 0.333930 | nan | -2.222108 | nan | nan | nan | 0.120233 | nan | nan | nan | -0.585597 | nan | nan | -1.454731 | nan | 0.470726 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.955375 | -1.255837 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.372987 | nan | nan | 1.995031 | nan | nan | nan | -0.188035 | nan | nan | -0.179925 | nan | nan | nan | nan | nan | nan | -0.340091 | nan | nan | -0.871518 | 1.115365 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.867050 | nan | nan | nan | -1.056383 | nan | nan | 0.412622 | 0.198457 | nan | nan | nan | nan | nan | nan | nan | -1.028291 | -0.761279 | nan | nan | nan | nan | nan | nan | nan | 0.779121 | nan | nan | nan | nan | nan | nan | -0.633336 | -0.127695 | nan | nan | nan | nan | nan | nan | nan | 0.175828 | nan | nan | nan | nan | nan | nan | nan | 0.220810 | 1.906694 | -0.769774 | 1.141544 | nan | nan | 0.148053 | nan | nan | -0.493245 | nan | nan | nan | nan | -1.279935 | 0.793184 | nan | nan | nan | -0.408281 | 0.572545 | -0.335814 | nan | nan | -0.278947 | nan | -1.045371 | nan | nan | nan | 0.655827 | nan | -0.839004 | nan | -0.593631 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.212700 | -1.616937 | nan | nan | nan | 0.252150 | nan | -0.421107 | nan | -1.389338 | nan | nan | nan | nan | nan | nan | nan | -1.094746 | nan | nan | nan | nan | nan | nan | -0.881483 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 3.894794 | 1.396125 | nan | nan | nan | 1.640365 | nan | nan | nan | nan | -1.230901 | nan | nan | nan | 1.346605 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.082465 | nan | nan | -0.710069 | nan | -1.089082 | nan | -1.185716 | nan | nan | nan | nan | nan | 1.357356 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.179706 | 0.778520 | 0.790346 | nan | nan | nan | 0.492500 | -0.722607 | nan | nan | 0.607282 | nan | nan | nan | -1.436072 | 0.068759 | nan | nan | nan | nan | nan | nan | -1.488441 | 1.208220 | nan | nan | -0.586152 | nan | nan | nan | nan | -0.463199 | nan | nan | nan | nan | nan | 0.490009 | -1.627624 | -0.364351 | nan | nan | nan | nan | -0.514077 | nan | nan | nan | nan | nan | nan | nan | -0.204468 | nan | nan | nan | nan | nan | nan | nan | nan | 0.658498 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.091533 | -1.166208 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.626637 | nan | nan | -1.109417 | nan | nan | nan | nan | nan | -0.388218 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.455506 | nan | nan | 0.443298 | -0.000956 | -1.520597 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.171781 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.156926 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.419963 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.225532 | nan | 0.185515 | nan | nan | nan | nan | nan | nan | -1.111053 | nan | 0.759797 | nan | nan | -0.801270 | nan | -0.285606 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | AADSIPSISPTGIITPTPTQSGMVSNCNKFYDVHSNDGCSAIASSQHVDLSSLYKWNPAIKTDCSGLQASVYVCIGILTTSMSTTSKLPTTTSKPSTTSKPPTGITTPTPTQSGMVKNCNKFYDVHSGDGCSAIASSQKVNLSSFYLWNPAVKTDCSGLQASVFVCIGTLTTSMSTTTSKLPTTTSKPSTTSKPPTGIITPTPTQNGMVKNCNKFYDVHSGDGCSAIASSQKVNLSSFYLWNPAVKTDCSGLQASVFVCVGTATTTTAAGITTPTPTQSGMVSGCNKFYDVHAGDGCSAIASSQKIALSSLYKWNPAVKTDCSGLQASVYICIGVGNAAAARVTGALEQKLISEEDLNSAVDHHHHHH | TAL6-4LysM |
| 1 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.210341 | -0.317977 | nan | -0.535696 | -0.428562 | 0.304758 | -0.599696 | nan | nan | -0.514044 | nan | -0.621800 | 0.110234 | 1.704315 | nan | -0.342968 | nan | nan | nan | nan | nan | nan | -0.241699 | -0.094321 | nan | nan | nan | 0.548443 | nan | -0.164111 | -0.301651 | nan | -0.286659 | nan | 5.996096 | nan | 0.195205 | nan | nan | nan | nan | nan | -0.572861 | -0.078468 | nan | nan | nan | -0.271587 | -0.185770 | -0.477049 | -0.148226 | 0.273879 | 3.048219 | nan | 1.536058 | nan | nan | nan | nan | nan | -0.388298 | 0.720448 | nan | -0.154050 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.145095 | nan | nan | nan | -0.120509 | -0.642615 | nan | nan | nan | nan | nan | nan | -0.358526 | nan | nan | nan | -0.401998 | nan | nan | nan | nan | -0.453027 | -0.597313 | 0.746013 | nan | 0.881687 | nan | nan | nan | nan | nan | nan | -0.673569 | -0.409280 | nan | nan | nan | -0.241929 | nan | nan | nan | nan | nan | nan | -0.084825 | nan | nan | nan | nan | -0.452808 | -0.526735 | 0.338334 | -0.350185 | -0.280775 | nan | nan | -0.219135 | nan | -0.214087 | nan | -0.278396 | nan | -0.039948 | 0.319400 | 0.767513 | nan | nan | 0.321303 | 0.054521 | nan | -0.004854 | -0.394323 | -0.397670 | -0.363056 | 0.408759 | nan | nan | nan | nan | -0.538161 | nan | -0.072649 | -0.305646 | -0.598274 | -0.146399 | -0.588251 | nan | 0.127787 | nan | nan | nan | -0.389585 | nan | nan | nan | -0.544289 | nan | -0.273656 | nan | 0.496878 | -0.562748 | 0.696372 | nan | nan | nan | 0.292971 | -0.439380 | nan | nan | -0.487665 | -0.387022 | -0.723142 | -0.716450 | 1.611207 | nan | nan | nan | -0.422152 | nan | 0.319929 | nan | -0.606341 | -0.160275 | -0.494404 | nan | nan | 0.049719 | 1.054716 | nan | -1.226240 | -0.689861 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.394123 | nan | nan | nan | nan | nan | nan | nan | nan | 0.209877 | -0.632032 | -0.279757 | nan | 0.120900 | -0.035907 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.393659 | nan | nan | nan | nan | nan | nan | nan | -0.163429 | nan | nan | nan | -0.735791 | nan | nan | nan | nan | -0.050865 | nan | nan | nan | nan | nan | nan | nan | nan | 0.153107 | nan | nan | nan | nan | nan | 0.114519 | -0.238581 | -0.506776 | 0.084050 | nan | nan | -0.195494 | nan | nan | nan | -0.169214 | nan | nan | nan | nan | nan | -0.537188 | -0.664153 | nan | nan | nan | 0.108785 | nan | nan | nan | nan | -0.017564 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.278367 | nan | nan | -0.706053 | nan | -0.421375 | -0.637758 | 0.325945 | -0.185604 | -0.206844 | -1.225292 | nan | -0.201082 | -0.284139 | -0.316140 | nan | 1.195916 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.161080 | nan | nan | nan | nan | nan | -0.551468 | nan | nan | nan | -0.120453 | 3.751031 | nan | nan | nan | -0.044979 | nan | nan | nan | nan | nan | -0.627522 | nan | nan | nan | nan | nan | 0.714630 | nan | nan | -0.307018 | -0.428596 | 3.366465 | nan | -0.475412 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.156273 | nan | -0.273549 | nan | nan | nan | nan | nan | nan | -0.257248 | nan | nan | nan | nan | nan | nan | -0.077096 | nan | nan | nan | nan | -0.238332 | nan | nan | -0.229461 | nan | 0.020193 | -0.151334 | nan | -0.297753 | nan | -0.169976 | nan | nan | nan | nan | -0.071467 | nan | nan | nan | nan | -0.118106 | nan | nan | nan | nan | nan | nan | nan | -0.315901 | nan | -0.184823 | nan | nan | nan | 0.176021 | -0.056910 | -0.571056 | 0.493649 | 0.506835 | 1.096384 | nan | nan | nan | nan | nan | -0.285038 | nan | -0.328707 | 0.059036 | 1.714136 | -0.023349 | nan | nan | 1.438082 | nan | nan | nan | nan | 0.560742 | nan | nan | -0.064961 | nan | nan | nan | 0.293321 | nan | -0.738018 | 0.159939 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.438784 | -0.647943 | -0.911864 | nan | -0.147095 | nan | nan | nan | nan | nan | nan | nan | -0.076763 | -0.105693 | nan | -0.291598 | nan | -0.332561 | nan | nan | nan | -0.471061 | nan | nan | nan | nan | nan | -0.623435 | nan | nan | nan | -0.423218 | nan | 0.335238 | nan | nan | nan | nan | nan | nan | nan | nan | -0.458750 | nan | -0.458470 | nan | nan | -0.620907 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.433047 | -0.746319 | 0.103050 | nan | nan | -0.424069 | -0.219640 | nan | 3.941640 | nan | -0.204654 | 0.140085 | nan | nan | 0.004276 | nan | 0.558532 | 0.196721 | -0.267338 | -0.263195 | nan | 2.066318 | 3.411378 | -0.375306 | -0.932223 | nan | nan | nan | nan | -0.189840 | nan | nan | nan | nan | -0.193638 | -0.518012 | nan | nan | nan | nan | nan | nan | -0.587022 | nan | nan | nan | nan | nan | nan | nan | 0.464959 | nan | 0.522682 | 2.988462 | nan | 1.023056 | 0.986010 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.209109 | -0.503419 | 0.011710 | -0.336139 | 5.985190 | 1.319522 | nan | 1.268927 | 2.710962 | nan | nan | nan | 0.528672 | nan | nan | nan | nan | 4.452175 | 1.298506 | 4.112301 | nan | nan | nan | nan | nan | nan | 2.141467 | 1.655698 | 4.640982 | nan | nan | nan | 0.770460 | 0.433840 | 3.938218 | 2.332191 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.565793 | nan | nan | 2.655383 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.949440 | nan | nan | nan | nan | nan | nan | nan | -0.135356 | nan | nan | nan | nan | nan | nan | -0.320277 | 0.660683 | nan | nan | nan | nan | -0.071170 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.039727 | nan | 0.091177 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.504007 | nan | nan | -0.307391 | nan | -0.085958 | nan | nan | nan | nan | nan | -1.266585 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.148673 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.367992 | nan | nan | nan | nan | nan | nan | -0.308856 | nan | nan | -0.266432 | nan | nan | nan | nan | -0.303892 | -0.416294 | -0.132050 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.316894 | -0.323859 | -0.596264 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.376123 | nan | -0.281852 | nan | nan | nan | nan | nan | -0.326506 | nan | nan | nan | -0.777421 | -0.576470 | nan | nan | nan | -0.662114 | -0.504215 | -0.713073 | nan | nan | nan | nan | -0.449993 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.243328 | nan | nan | nan | nan | nan | -0.628874 | nan | nan | nan | -0.574322 | -0.006513 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.048031 | nan | nan | 0.014492 | -0.096467 | 0.329244 | nan | nan | -0.444026 | nan | -0.496267 | nan | nan | nan | nan | nan | 0.571160 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.019903 | 0.037512 | nan | 3.418474 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.062730 | nan | nan | nan | -0.072516 | nan | nan | -0.267884 | nan | nan | nan | nan | nan | nan | nan | nan | -0.340286 | nan | -0.693033 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.578903 | nan | nan | nan | nan | nan | nan | -0.260458 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.029600 | nan | 0.085726 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.061387 | -0.230035 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.068152 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.146414 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.929766 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.021350 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.188295 | nan | nan | nan | nan | nan | nan | -0.613549 | nan | nan | nan | nan | nan | nan | nan | -0.792706 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.305427 | nan | nan | nan | nan | nan | nan | nan | -0.191716 | nan | -0.786313 | -0.728967 | -1.106288 | -0.380770 | -0.522477 | 0.091364 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.294415 | nan | nan | -0.603658 | nan | nan | nan | nan | nan | nan | -0.374202 | nan | nan | 1.311709 | -0.318125 | -0.164150 | nan | nan | nan | nan | -0.226948 | 1.353726 | -0.425933 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.031533 | nan | -0.276207 | nan | nan | nan | nan | nan | -0.196006 | -0.642186 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.004902 | nan | -0.618264 | -0.116827 | nan | -0.626297 | -0.454391 | 0.076760 | -0.352582 | -0.546664 | nan | -0.255336 | nan | nan | nan | 3.022814 | 0.405144 | -0.859338 | nan | nan | nan | -0.214939 | 0.698437 | -1.061306 | nan | -0.268759 | 0.053256 | -0.432154 | nan | -0.453639 | nan | nan | nan | -0.132935 | nan | nan | 0.249749 | -0.117152 | -0.124793 | -0.667753 | nan | nan | nan | nan | nan | nan | -0.560969 | nan | 0.734671 | nan | nan | nan | -0.363380 | nan | nan | -0.723359 | nan | -0.541242 | nan | nan | nan | nan | nan | nan | nan | 0.711890 | nan | nan | -0.274278 | -0.208130 | nan | -0.146078 | -0.305149 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.002285 | -0.312072 | nan | -0.194684 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.135019 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.192462 | nan | nan | nan | -0.880676 | nan | 0.222977 | -0.261812 | nan | nan | 0.026415 | nan | nan | nan | nan | nan | nan | nan | -0.099068 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.432871 | nan | nan | nan | nan | -0.570354 | 0.842646 | nan | nan | nan | nan | nan | -0.958361 | 0.133986 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.006577 | nan | nan | nan | nan | -0.482137 | nan | nan | nan | nan | nan | nan | -0.986270 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.718850 | nan | nan | nan | nan | nan | nan | nan | nan | -0.650632 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.022822 | nan | nan | nan | nan | nan | nan | nan | nan | -0.085626 | 0.139780 | nan | nan | nan | -0.211927 | nan | nan | -0.498004 | nan | nan | nan | nan | nan | -0.763792 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.160861 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.247478 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.334203 | nan | nan | nan | -0.476253 | nan | nan | -0.224135 | nan | nan | nan | -0.333129 | 0.303401 | -0.283198 | nan | nan | nan | nan | -0.334101 | -0.094590 | nan | 5.370085 | -0.467862 | 0.155618 | nan | nan | nan | 0.092594 | nan | -0.252477 | nan | -0.121639 | nan | nan | 0.011680 | 0.200834 | 0.753740 | nan | nan | -0.549464 | -0.194579 | nan | -0.238250 | nan | -0.348041 | -0.474801 | -0.429747 | nan | nan | nan | nan | nan | nan | nan | 0.044244 | nan | nan | nan | 0.341388 | -0.728712 | 0.283256 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.329709 | 0.096213 | -0.431491 | nan | nan | -0.417758 | nan | nan | nan | -0.712527 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.042465 | -0.115827 | nan | nan | nan | nan | nan | nan | -0.931021 | nan | -0.432294 | nan | nan | nan | nan | -0.022245 | nan | nan | 1.624166 | nan | nan | nan | nan | nan | -0.539692 | nan | nan | nan | nan | nan | nan | nan | 4.099614 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.397088 | 0.489274 | -0.373275 | -0.136173 | nan | nan | nan | -0.263554 | nan | -0.192120 | nan | nan | nan | -0.107791 | nan | nan | nan | 0.089840 | nan | nan | -0.210930 | nan | -0.489136 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.535536 | -1.350794 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.441527 | nan | nan | 0.271697 | nan | nan | nan | 0.143828 | nan | nan | 0.502451 | nan | nan | nan | nan | nan | nan | -0.389743 | nan | nan | 0.160502 | -0.443902 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.759362 | nan | nan | nan | -0.672274 | nan | nan | -0.763808 | 0.043080 | nan | nan | nan | nan | nan | nan | nan | -0.324763 | -0.735311 | nan | nan | nan | nan | nan | nan | nan | 0.606204 | nan | nan | nan | nan | nan | nan | -0.142990 | -0.537923 | nan | nan | nan | nan | nan | nan | nan | 0.024373 | nan | nan | nan | nan | nan | nan | nan | -0.232817 | -0.253269 | -0.167730 | -0.302303 | nan | nan | -0.272344 | nan | nan | -0.465127 | nan | nan | nan | nan | -0.337413 | -0.337699 | nan | nan | nan | -0.048925 | -0.204174 | -0.331235 | nan | nan | -0.275976 | nan | -0.112222 | nan | nan | nan | -0.372519 | nan | -0.381540 | nan | -0.629240 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.212922 | -0.068444 | nan | nan | nan | 0.274100 | nan | -0.269315 | nan | -0.020473 | nan | nan | nan | nan | nan | nan | nan | 0.059676 | nan | nan | nan | nan | nan | nan | 3.188752 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.179521 | 2.341252 | nan | nan | nan | -0.716474 | nan | nan | nan | nan | -0.502049 | nan | nan | nan | 0.044380 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.190638 | nan | nan | 0.066739 | nan | -0.347539 | nan | -0.387395 | nan | nan | nan | nan | nan | -0.577035 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.379801 | -0.632533 | -0.457350 | nan | nan | nan | -0.601558 | -0.895900 | nan | nan | -0.141286 | nan | nan | nan | -0.272919 | 0.387567 | nan | nan | nan | nan | nan | nan | -0.526855 | -0.847587 | nan | nan | -0.384928 | nan | nan | nan | nan | -0.715208 | nan | nan | nan | nan | nan | -0.552275 | -0.435920 | 0.673410 | nan | nan | nan | nan | -0.806533 | nan | nan | nan | nan | nan | nan | nan | -0.316786 | nan | nan | nan | nan | nan | nan | nan | nan | -0.765163 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.665418 | -0.547333 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.188309 | nan | nan | 2.013600 | nan | nan | nan | nan | nan | -0.285953 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.114993 | nan | nan | -0.318728 | -0.609542 | -0.444485 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.288337 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.236235 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.087587 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.230830 | nan | -0.591006 | nan | nan | nan | nan | nan | nan | -0.542732 | nan | -0.645822 | nan | nan | -0.630642 | nan | -0.083163 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | AAFFSLVVLLALLPFGIHASALPSTELTPRVNPNLPGPNDVFVGFRGTNNIVGAIYEKDGIKDTPLNNGADAEMGPGSYITDGLFTARIFASGSAGNNPGTTATVCKIYASGVQWRGAPKGYIPVAYFGNGPTAEQRRQTYLRTLGLPGTAIRFGNLNTNPGQGPTAPRQMVVPPGPLRSLLKASCFPAADTRLPQTLHYFEKRVDWNIITAADSQTWGH | rCnSL-proA |
| 2 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.141335 | -0.048689 | nan | 0.028283 | -1.017600 | -0.871024 | -1.540761 | nan | nan | -0.350449 | nan | -0.652191 | 0.716365 | 0.612228 | nan | -1.258691 | nan | nan | nan | nan | nan | nan | 0.237488 | 0.422113 | nan | nan | nan | -1.543964 | nan | -1.187212 | 0.457266 | nan | 0.480527 | nan | 0.420569 | nan | 0.614918 | nan | nan | nan | nan | nan | -0.839644 | -0.770383 | nan | nan | nan | -0.064917 | 1.149108 | -0.565169 | 1.387117 | 1.495458 | 1.109946 | nan | 0.655759 | nan | nan | nan | nan | nan | -1.040241 | 0.724848 | nan | 0.654293 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.034218 | nan | nan | nan | 0.338654 | -0.844789 | nan | nan | nan | nan | nan | nan | 1.889215 | nan | nan | nan | -0.715226 | nan | nan | nan | nan | 0.076582 | 4.683932 | 2.555725 | nan | 0.208505 | nan | nan | nan | nan | nan | nan | 0.169653 | 1.500775 | nan | nan | nan | 0.835543 | nan | nan | nan | nan | nan | nan | -0.889909 | nan | nan | nan | nan | -1.078300 | 0.470129 | -0.124160 | -0.552058 | 0.254316 | nan | nan | -0.227861 | nan | 0.123434 | nan | -0.957585 | nan | -0.863318 | -0.497218 | -0.592302 | nan | nan | 1.597832 | 0.337272 | nan | -0.571593 | -1.002809 | -0.527701 | 0.173993 | 3.104163 | nan | nan | nan | nan | -0.213374 | nan | -0.495262 | -0.016564 | 0.825037 | -0.161087 | -0.145769 | nan | -0.200245 | nan | nan | nan | -1.128504 | nan | nan | nan | -0.354246 | nan | -0.495097 | nan | 0.349565 | 1.033642 | -0.775093 | nan | nan | nan | 0.738992 | -0.331531 | nan | nan | -0.729900 | -0.846219 | -0.980832 | 1.013321 | 0.155313 | nan | nan | nan | -0.715678 | nan | 0.176707 | nan | 0.387436 | 0.200393 | -0.876682 | nan | nan | 0.390241 | 1.570174 | nan | 0.887831 | 0.107227 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.171433 | nan | nan | nan | nan | nan | nan | nan | nan | -0.335483 | 0.825236 | -0.374665 | nan | 0.125660 | 0.760958 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.626154 | nan | nan | nan | nan | nan | nan | nan | 0.906766 | nan | nan | nan | -0.372149 | nan | nan | nan | nan | 2.642953 | nan | nan | nan | nan | nan | nan | nan | nan | 0.145219 | nan | nan | nan | nan | nan | 0.209840 | 0.537053 | 3.679210 | 0.472882 | nan | nan | 0.837446 | nan | nan | nan | 0.931615 | nan | nan | nan | nan | nan | -0.390090 | -0.559610 | nan | nan | nan | -0.388097 | nan | nan | nan | nan | -0.590225 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.595967 | nan | nan | -0.196759 | nan | -0.563083 | -0.640716 | 3.466913 | 2.789688 | 0.992895 | 0.369458 | nan | -0.763190 | -0.982501 | -0.101580 | nan | 1.548296 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.152511 | nan | nan | nan | nan | nan | -0.128297 | nan | nan | nan | -1.036687 | 0.993182 | nan | nan | nan | 1.360269 | nan | nan | nan | nan | nan | 0.240559 | nan | nan | nan | nan | nan | 2.055367 | nan | nan | 0.900653 | -0.269334 | -0.010582 | nan | -0.067021 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.879200 | nan | -0.190946 | nan | nan | nan | nan | nan | nan | -0.551266 | nan | nan | nan | nan | nan | nan | 0.084391 | nan | nan | nan | nan | -0.051376 | nan | nan | -0.409344 | nan | 1.010344 | 0.077779 | nan | -0.620924 | nan | 0.437636 | nan | nan | nan | nan | -0.798348 | nan | nan | nan | nan | -0.731754 | nan | nan | nan | nan | nan | nan | nan | 0.211020 | nan | -0.673579 | nan | nan | nan | 0.451430 | -0.152162 | 0.099131 | 0.484088 | 1.129303 | 0.815293 | nan | nan | nan | nan | nan | 1.591959 | nan | -0.068063 | 0.484870 | -0.071728 | 1.148531 | nan | nan | 5.611608 | nan | nan | nan | nan | -1.076119 | nan | nan | -0.401547 | nan | nan | nan | -0.768734 | nan | -0.215077 | -1.157278 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.879382 | -1.147433 | -0.090986 | nan | 0.093784 | nan | nan | nan | nan | nan | nan | nan | -0.532454 | 0.064582 | nan | 0.264495 | nan | -1.390338 | nan | nan | nan | 0.182229 | nan | nan | nan | nan | nan | 3.096448 | nan | nan | nan | 0.648753 | nan | -0.774947 | nan | nan | nan | nan | nan | nan | nan | nan | -0.670909 | nan | -1.083155 | nan | nan | 0.658986 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.098134 | -1.495887 | 0.297815 | nan | nan | 1.203526 | -0.744118 | nan | 1.048740 | nan | 0.599308 | -0.225759 | nan | nan | -1.123019 | nan | 1.807711 | -0.213169 | -0.554981 | -1.220352 | nan | -0.417603 | -0.250864 | -0.581462 | -0.072945 | nan | nan | nan | nan | 1.673273 | nan | nan | nan | nan | 0.216192 | -0.498000 | nan | nan | nan | nan | nan | nan | 0.472995 | nan | nan | nan | nan | nan | nan | nan | -0.518564 | nan | -0.310421 | 0.027248 | nan | 0.598330 | -0.573974 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.492076 | 0.173055 | -0.380729 | -0.087446 | 0.700929 | 2.037591 | nan | 1.555403 | 1.535190 | nan | nan | nan | -0.395869 | nan | nan | nan | nan | 0.926895 | 1.394170 | 1.808713 | nan | nan | nan | nan | nan | nan | 2.517612 | 3.269472 | 2.270964 | nan | nan | nan | 2.197590 | 1.300799 | 1.714303 | 3.905954 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.995075 | nan | nan | 0.339238 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.398401 | nan | nan | nan | nan | nan | nan | nan | -0.717286 | nan | nan | nan | nan | nan | nan | -1.151917 | -0.738147 | nan | nan | nan | nan | 0.600467 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.429460 | nan | 0.525111 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.192369 | nan | nan | -0.203349 | nan | 0.419092 | nan | nan | nan | nan | nan | -0.923743 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.487743 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.051212 | nan | nan | nan | nan | nan | nan | -0.670214 | nan | nan | 0.514861 | nan | nan | nan | nan | 0.494058 | -0.822357 | 0.443901 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.164600 | 0.603060 | 0.435522 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.506143 | nan | -1.142534 | nan | nan | nan | nan | nan | -1.278513 | nan | nan | nan | -0.028601 | 0.363345 | nan | nan | nan | -0.109694 | 1.009996 | 0.781620 | nan | nan | nan | nan | 0.522848 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.000915 | nan | nan | nan | nan | nan | 0.326645 | -0.515516 | nan | nan | -0.223894 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.411156 | nan | nan | 0.730003 | -0.843211 | -1.058670 | nan | nan | 1.370434 | nan | -0.475073 | nan | nan | nan | nan | nan | -0.825378 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.185191 | -0.796588 | nan | -0.867760 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.774234 | nan | nan | nan | -1.301059 | nan | nan | -0.454536 | nan | nan | nan | nan | nan | nan | nan | nan | -1.462383 | nan | -0.649293 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.462466 | nan | nan | nan | nan | nan | nan | 0.953470 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.184253 | nan | -1.188183 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.587275 | 0.176616 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.226841 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.364115 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.100772 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.575073 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.668801 | nan | nan | nan | nan | nan | nan | -1.077393 | nan | nan | nan | nan | nan | nan | nan | -0.668994 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.723625 | nan | nan | nan | nan | nan | nan | nan | -0.217599 | nan | -0.792700 | -0.916332 | 0.081845 | 1.468745 | -0.892370 | 0.069646 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.177320 | nan | nan | -1.058256 | nan | nan | nan | nan | nan | nan | -0.622087 | nan | nan | -0.727465 | -1.108628 | 1.460117 | nan | nan | nan | nan | 0.082347 | 0.953946 | -0.813898 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.122677 | nan | -1.349787 | nan | nan | nan | nan | nan | -0.894629 | -0.588259 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.017165 | nan | -0.153106 | -0.295210 | nan | -1.245724 | 0.116423 | -0.320367 | 0.424547 | 0.978872 | nan | -0.330253 | nan | nan | nan | 2.186425 | 0.477924 | -0.456296 | nan | nan | nan | -0.017359 | 0.510488 | 1.956776 | nan | 1.126133 | 3.226699 | 0.602039 | nan | -1.005354 | nan | nan | nan | -0.624235 | nan | nan | -0.581631 | -0.907495 | 0.271080 | -0.012440 | nan | nan | nan | nan | nan | nan | -0.709801 | nan | -1.029425 | nan | nan | nan | -0.953545 | nan | nan | -1.174501 | nan | -0.557060 | nan | nan | nan | nan | nan | nan | nan | -0.561542 | nan | nan | 0.034974 | 0.168207 | nan | 0.051790 | -0.579770 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.642614 | -0.001472 | nan | -0.654685 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.769802 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.902390 | nan | nan | nan | 0.431743 | nan | 0.137663 | -0.618213 | nan | nan | 1.133564 | nan | nan | nan | nan | nan | nan | nan | -1.054152 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.160963 | nan | nan | nan | nan | -1.512611 | -0.708850 | nan | nan | nan | nan | nan | -1.480671 | 0.974897 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.221059 | nan | nan | nan | nan | 0.520033 | nan | nan | nan | nan | nan | nan | -0.805138 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.367533 | nan | nan | nan | nan | nan | nan | nan | nan | -0.990863 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.130424 | nan | nan | nan | nan | nan | nan | nan | nan | -0.420475 | -0.421585 | nan | nan | nan | -0.410854 | nan | nan | -0.692633 | nan | nan | nan | nan | nan | -0.277207 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.516160 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.740449 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.346629 | nan | nan | nan | -0.763731 | nan | nan | 0.491866 | nan | nan | nan | -0.369327 | 0.854738 | -0.385137 | nan | nan | nan | nan | -0.142725 | -0.920891 | nan | 0.302544 | -0.192656 | nan | nan | nan | nan | -0.515233 | nan | -1.160781 | nan | -0.806743 | nan | nan | -0.890371 | 0.015838 | -1.254571 | nan | nan | -0.416332 | -0.256326 | nan | -0.386644 | nan | 0.442694 | 0.015878 | -1.056443 | nan | nan | nan | nan | nan | nan | nan | -0.099207 | nan | nan | nan | -0.467975 | 0.012557 | 0.920586 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.300478 | -1.077420 | -0.724094 | 0.406078 | nan | -1.348789 | nan | nan | nan | 0.080169 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.009904 | -0.940578 | nan | nan | nan | nan | nan | nan | -1.178166 | nan | 0.366113 | nan | nan | nan | nan | -0.615617 | nan | nan | -0.057792 | nan | nan | nan | nan | nan | -0.964989 | nan | nan | nan | nan | nan | nan | nan | 1.396847 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.421386 | 0.323981 | -0.290309 | -0.593175 | nan | nan | nan | -1.272687 | nan | 0.764037 | nan | nan | nan | -1.013082 | nan | nan | nan | -0.572224 | nan | nan | 0.144831 | nan | -0.627496 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.523728 | 2.242855 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.529255 | nan | nan | -0.000575 | nan | nan | nan | -0.070429 | nan | nan | 1.568826 | nan | nan | nan | nan | nan | nan | 2.321792 | nan | nan | 0.167485 | -1.036465 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.680242 | nan | nan | nan | -0.675306 | nan | nan | -1.048313 | -0.789362 | nan | nan | nan | nan | nan | nan | nan | -0.635699 | -0.450291 | nan | nan | nan | nan | nan | nan | nan | 0.843699 | nan | nan | nan | nan | nan | nan | 0.222346 | -1.554352 | nan | nan | nan | nan | nan | nan | nan | 0.439390 | nan | nan | nan | nan | nan | nan | nan | -0.114403 | -0.451667 | -0.521770 | -0.244926 | nan | nan | 0.156868 | nan | nan | -0.928180 | nan | nan | nan | nan | 0.483767 | 0.381458 | nan | nan | nan | -0.329480 | 1.668495 | -0.618409 | nan | nan | 0.992757 | nan | -0.280346 | nan | nan | nan | -1.050066 | nan | -0.891082 | nan | -1.133454 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.285970 | 1.051619 | nan | nan | nan | -1.100358 | nan | -0.213941 | nan | -0.823419 | nan | nan | nan | nan | nan | nan | nan | -1.168836 | nan | nan | nan | nan | nan | nan | 1.737089 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.349192 | 0.793185 | nan | nan | nan | 0.305781 | nan | nan | nan | nan | 0.246530 | nan | nan | nan | 0.206354 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.080515 | nan | nan | 0.255455 | nan | -0.279725 | nan | -1.322618 | nan | nan | nan | nan | nan | 1.060777 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.370770 | -1.167221 | -0.769847 | nan | nan | nan | -0.597977 | -0.712772 | nan | nan | -0.380666 | nan | nan | nan | 0.241274 | -0.035084 | nan | nan | nan | nan | nan | nan | -0.254492 | -0.756890 | nan | nan | -0.073090 | nan | nan | nan | nan | 0.406664 | nan | nan | nan | nan | nan | -0.735348 | -0.756698 | 0.119479 | nan | nan | nan | nan | 0.774519 | nan | nan | nan | nan | nan | nan | nan | 0.749350 | nan | nan | nan | nan | nan | nan | nan | nan | -0.976446 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.331058 | -1.136037 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.544240 | nan | nan | -1.291898 | nan | nan | nan | nan | nan | -0.505057 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.716022 | nan | nan | -0.884804 | -0.071245 | -0.464590 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.883028 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.707221 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.797868 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.493683 | nan | 1.662689 | nan | nan | nan | nan | nan | nan | -0.703489 | nan | -1.168415 | nan | nan | -1.297091 | nan | 1.005849 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | AANEADYQAKLTAYQTELARVQKANADAKAAYEAAVAANNAANAALTAENTAIKKRNADAKADYEAKLAKYQADLAKYQKDLADYPVKLKAYEDEQASIKAALAELEKHKNEDGNLTEPSAQNLVYDLEPNANLSLTTDGKFLKASAVDDAFSKSTSKAKYDQKILQLDDLDITNLEQSNDVASSMELYGNFGDKAGWSTTVSNNSQVKWGSVLLERGQSATATYTNLQNSYYNGKKISKIVYKYTVDPKSKFQGQKVWLGIFTDPTLGVFASAYTGQVEKNTSIFIKNEFTFYDEDGKPINFDNALLSVASLNREHNSIEMAKDYSGKFVKISGSSIGEKNGMIYATDTLNFKQGEGGSRWTMYKNSQAGSGWDSSDAPNSWYGAGAIKMSGPNNYVTVGATSATNVMPVSDMPVVPGKDNTDGKKPNIWYSLNGKIRAVNVPKVTKEKPTPPVKPTAPTKPTYETEKPLKPAPVAPNYEKEPTPPTRLEHHHHHH | AntigenI/IIA3VP1 |
| 3 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.951637 | -0.590611 | nan | -0.209295 | -1.768618 | 0.072966 | -0.120822 | nan | nan | -0.588345 | nan | -0.073485 | -1.229778 | -0.903323 | nan | -0.233609 | nan | nan | nan | nan | nan | nan | -0.595171 | 0.213082 | nan | nan | nan | 0.974062 | nan | -1.468367 | 0.905727 | nan | -1.174894 | nan | -2.406395 | nan | 0.448905 | nan | nan | nan | nan | nan | -0.674998 | -0.124440 | nan | nan | nan | 0.310361 | 0.072212 | 1.268628 | -0.376546 | 0.429747 | -1.153319 | nan | 0.463288 | nan | nan | nan | nan | nan | -0.313873 | -1.396695 | nan | 0.083592 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.064160 | nan | nan | nan | -0.205204 | 0.428165 | nan | nan | nan | nan | nan | nan | 3.118173 | nan | nan | nan | 0.291483 | nan | nan | nan | nan | -0.195754 | 0.066959 | 0.008242 | nan | -0.207449 | nan | nan | nan | nan | nan | nan | -0.762228 | 2.678329 | nan | nan | nan | 1.621970 | nan | nan | nan | nan | nan | nan | 1.821577 | nan | nan | nan | nan | 0.063804 | -0.395757 | 1.447813 | -0.384916 | 0.326316 | nan | nan | 0.612385 | nan | -0.805186 | nan | 0.209411 | nan | 1.378479 | -0.506181 | -1.232264 | nan | nan | -0.132982 | 0.722559 | nan | 0.490832 | 1.196810 | -0.907167 | 0.478301 | 1.704462 | nan | nan | nan | nan | -0.999434 | nan | -0.723832 | -0.509363 | -0.694436 | -0.889037 | 0.955400 | nan | 0.687000 | nan | nan | nan | -0.756971 | nan | nan | nan | -0.872953 | nan | -0.404418 | nan | 0.346657 | -0.391230 | -1.064233 | nan | nan | nan | 0.203522 | 1.125154 | nan | nan | -0.723746 | 0.557047 | 0.179450 | 0.420203 | -2.017785 | nan | nan | nan | -0.762109 | nan | -0.144362 | nan | -0.337741 | 1.582901 | 1.535322 | nan | nan | 0.713354 | -0.108216 | nan | -0.239374 | -0.198454 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.348999 | nan | nan | nan | nan | nan | nan | nan | nan | 0.625678 | -0.409209 | 0.367381 | nan | -0.825724 | 0.672665 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.406999 | nan | nan | nan | nan | nan | nan | nan | -0.013477 | nan | nan | nan | -0.382688 | nan | nan | nan | nan | 0.693050 | nan | nan | nan | nan | nan | nan | nan | nan | -0.638029 | nan | nan | nan | nan | nan | 0.506819 | -0.760823 | 2.958841 | -0.229898 | nan | nan | 0.160223 | nan | nan | nan | 2.991649 | nan | nan | nan | nan | nan | 0.001304 | -0.003395 | nan | nan | nan | -0.873890 | nan | nan | nan | nan | -0.452589 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.943522 | nan | nan | -0.439595 | nan | -0.376261 | 0.127424 | -0.832081 | 0.586475 | -1.257440 | 1.501641 | nan | -0.260007 | 2.223311 | 0.538368 | nan | 0.080427 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.018306 | nan | nan | nan | nan | nan | -0.541649 | nan | nan | nan | -0.417671 | -0.687471 | nan | nan | nan | 0.186877 | nan | nan | nan | nan | nan | 0.464817 | nan | nan | nan | nan | nan | -0.972353 | nan | nan | 2.693062 | 0.404590 | -0.586090 | nan | -0.008127 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.603526 | nan | 0.524593 | nan | nan | nan | nan | nan | nan | -0.670450 | nan | nan | nan | nan | nan | nan | -0.191908 | nan | nan | nan | nan | 0.459009 | nan | nan | -0.343084 | nan | -2.145714 | -0.299270 | nan | -1.582209 | nan | -0.336853 | nan | nan | nan | nan | -0.503355 | nan | nan | nan | nan | -0.686884 | nan | nan | nan | nan | nan | nan | nan | -0.122241 | nan | 0.950281 | nan | nan | nan | -1.168348 | 3.656594 | 0.699332 | -0.231499 | -0.060755 | -0.157388 | nan | nan | nan | nan | nan | 1.473764 | nan | -0.601700 | -0.570523 | 0.355733 | -0.913045 | nan | nan | 0.008199 | nan | nan | nan | nan | -1.311791 | nan | nan | -0.428372 | nan | nan | nan | -1.391807 | nan | -0.728623 | 0.048011 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.847592 | -0.120010 | -1.012224 | nan | -0.288547 | nan | nan | nan | nan | nan | nan | nan | 0.801761 | 0.148875 | nan | -0.760349 | nan | 0.447414 | nan | nan | nan | -1.195392 | nan | nan | nan | nan | nan | 1.020333 | nan | nan | nan | -0.669842 | nan | -0.094799 | nan | nan | nan | nan | nan | nan | nan | nan | -0.503495 | nan | 0.532867 | nan | nan | -0.396656 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.915846 | -1.312695 | 0.667610 | nan | nan | -1.078118 | -0.744122 | nan | -0.177299 | nan | 0.066577 | -0.341433 | nan | nan | 0.478667 | nan | -1.432792 | -1.239424 | 0.569050 | 0.551174 | nan | -0.309745 | -1.184158 | -0.993529 | -0.218031 | nan | nan | nan | nan | -1.586408 | nan | nan | nan | nan | -0.133160 | -0.648386 | nan | nan | nan | nan | nan | nan | 0.445212 | nan | nan | nan | nan | nan | nan | nan | 0.340408 | nan | -1.108098 | -0.570270 | nan | 1.237671 | 0.536947 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.573462 | -1.816202 | -0.431790 | -1.181353 | -0.047435 | 0.711572 | nan | 0.687726 | -0.076688 | nan | nan | nan | 0.635663 | nan | nan | nan | nan | -0.179234 | -0.033232 | 2.093702 | nan | nan | nan | nan | nan | nan | 1.773395 | -0.763503 | 0.045718 | nan | nan | nan | -1.114337 | 0.184397 | 0.687753 | 0.243150 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.432827 | nan | nan | 0.100575 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.642413 | nan | nan | nan | nan | nan | nan | nan | 0.867767 | nan | nan | nan | nan | nan | nan | 0.941375 | -0.192011 | nan | nan | nan | nan | 1.147492 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.093860 | nan | -1.597631 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.255916 | nan | nan | 2.004675 | nan | -0.644327 | nan | nan | nan | nan | nan | 0.256672 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.478630 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.856145 | nan | nan | nan | nan | nan | nan | 0.215771 | nan | nan | 0.692617 | nan | nan | nan | nan | -0.867506 | 1.857473 | 0.433973 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.547387 | 2.508914 | 2.997883 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.092527 | nan | 0.716433 | nan | nan | nan | nan | nan | -1.321049 | nan | nan | nan | -0.839434 | -1.524387 | nan | nan | nan | 0.457076 | 0.947794 | 0.000163 | nan | nan | nan | nan | -0.018150 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 2.440934 | nan | nan | nan | nan | nan | -1.350483 | nan | nan | nan | 0.453706 | -0.793529 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.284174 | nan | nan | -0.315127 | 0.375951 | -0.541687 | nan | nan | -0.839709 | nan | -0.240699 | nan | nan | nan | nan | nan | 0.272568 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.070937 | -1.474703 | nan | 1.817841 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.491648 | nan | nan | nan | 0.992186 | nan | nan | -0.302666 | nan | nan | nan | nan | nan | nan | nan | nan | -0.701536 | nan | 0.679663 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.076716 | nan | nan | nan | nan | nan | nan | 0.157181 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.578731 | nan | -0.352899 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.776132 | -0.389718 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.771739 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.128541 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.743590 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.114156 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.562752 | nan | nan | nan | nan | nan | nan | 2.024226 | nan | nan | nan | nan | nan | nan | nan | 0.145742 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.284101 | nan | nan | nan | nan | nan | nan | nan | 0.821275 | nan | -0.903356 | 0.444561 | -1.305724 | -1.485496 | -0.008224 | -0.364553 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.905344 | nan | nan | -0.296444 | nan | nan | nan | nan | nan | nan | 0.420031 | nan | nan | 0.127704 | -1.172369 | -0.838201 | nan | nan | nan | nan | -1.160263 | 1.142438 | -0.275461 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.558982 | nan | 1.148908 | nan | nan | nan | nan | nan | 0.222792 | 0.450762 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.653957 | nan | -0.361974 | 0.060989 | nan | 0.267777 | 0.020739 | -1.200785 | 0.822755 | 1.762667 | nan | 0.291381 | nan | nan | nan | 0.251733 | 0.195232 | -0.539022 | nan | nan | nan | 0.511997 | -0.349578 | -0.372837 | nan | -0.734474 | 1.317505 | 2.310868 | nan | -0.562195 | nan | nan | nan | 0.441697 | nan | nan | 0.159291 | -0.871995 | -0.152614 | -0.350767 | nan | nan | nan | nan | nan | nan | -0.097614 | nan | -0.043341 | nan | nan | nan | 1.903397 | nan | nan | -0.187887 | nan | -1.077811 | nan | nan | nan | nan | nan | nan | nan | -0.966938 | nan | nan | -0.196707 | 0.249518 | nan | 0.737324 | -0.323772 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.001472 | -1.185256 | nan | 1.316377 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.253259 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.312635 | nan | nan | nan | -1.176665 | nan | -0.558890 | 0.209981 | nan | nan | 1.879744 | nan | nan | nan | nan | nan | nan | nan | 2.709582 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.997819 | nan | nan | nan | nan | -0.014177 | -0.419786 | nan | nan | nan | nan | nan | 0.122311 | -0.099242 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.131879 | nan | nan | nan | nan | -0.128272 | nan | nan | nan | nan | nan | nan | -0.347904 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 2.524406 | nan | nan | nan | nan | nan | nan | nan | nan | -0.092632 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.021406 | nan | nan | nan | nan | nan | nan | nan | nan | -0.725861 | 0.135504 | nan | nan | nan | -0.273553 | nan | nan | 0.171989 | nan | nan | nan | nan | nan | -0.108189 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.529707 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.385468 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.154296 | nan | nan | nan | -1.434264 | nan | nan | -0.741321 | nan | nan | nan | -0.799795 | -0.659313 | 0.648770 | nan | nan | nan | nan | -0.539046 | 0.460526 | nan | -1.382269 | 0.806580 | nan | nan | nan | nan | 1.138331 | nan | -0.457176 | nan | 2.365913 | nan | nan | 0.273375 | -0.412471 | -0.031418 | nan | nan | 1.222739 | -0.755973 | nan | 1.023234 | nan | -1.626629 | -0.806674 | 0.886790 | nan | nan | nan | nan | nan | nan | nan | 0.865350 | nan | nan | nan | -0.572708 | -0.585570 | 0.220300 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.417197 | 0.049115 | 0.193585 | 0.484905 | nan | 0.012646 | nan | nan | nan | -0.682798 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.980696 | -0.720602 | nan | nan | nan | nan | nan | nan | -1.164252 | nan | -0.146854 | nan | nan | nan | nan | -0.540276 | nan | nan | -0.657763 | nan | nan | nan | nan | nan | 1.756741 | nan | nan | nan | nan | nan | nan | nan | -0.398303 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.926273 | -0.648273 | -0.274664 | 0.025699 | nan | nan | nan | -0.542532 | nan | -1.250211 | nan | nan | nan | 0.786932 | nan | nan | nan | -1.452149 | nan | nan | -1.140013 | nan | -0.459964 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.196312 | -0.793960 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.168743 | nan | nan | -0.591144 | nan | nan | nan | 0.057754 | nan | nan | -0.077485 | nan | nan | nan | nan | nan | nan | 0.957655 | nan | nan | -0.367007 | 0.058728 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.427600 | nan | nan | nan | -1.041480 | nan | nan | 0.267018 | -0.544599 | nan | nan | nan | nan | nan | nan | nan | -1.281566 | -1.083673 | nan | nan | nan | nan | nan | nan | nan | 1.671810 | nan | nan | nan | nan | nan | nan | 0.398731 | -0.907107 | nan | nan | nan | nan | nan | nan | nan | -0.730259 | nan | nan | nan | nan | nan | nan | nan | 0.029384 | -0.076599 | -0.567314 | -0.141536 | nan | nan | 0.695219 | nan | nan | 0.741668 | nan | nan | nan | nan | -0.593798 | -0.013122 | nan | nan | nan | 0.076119 | -0.890878 | 0.064321 | nan | nan | 0.005115 | nan | 0.838301 | nan | nan | nan | 0.106119 | nan | -0.010285 | nan | -1.406960 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.287860 | -0.828776 | nan | nan | nan | -1.095572 | nan | 1.521757 | nan | 1.366456 | nan | nan | nan | nan | nan | nan | nan | 1.285267 | nan | nan | nan | nan | nan | nan | 1.615747 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.316575 | 0.003150 | nan | nan | nan | -0.387150 | nan | nan | nan | nan | 2.128922 | nan | nan | nan | -1.094625 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.108200 | nan | nan | -0.092169 | nan | -1.033255 | nan | -0.007605 | nan | nan | nan | nan | nan | 0.283393 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.324903 | -0.721049 | 1.989544 | nan | nan | nan | -0.899679 | -1.003498 | nan | nan | -0.044499 | nan | nan | nan | -0.252589 | 0.051698 | nan | nan | nan | nan | nan | nan | -1.510983 | -1.090636 | nan | nan | -0.398503 | nan | nan | nan | nan | 2.166661 | nan | nan | nan | nan | nan | -0.138793 | -0.818538 | -0.053375 | nan | nan | nan | nan | 1.195357 | nan | nan | nan | nan | nan | nan | nan | 0.372549 | nan | nan | nan | nan | nan | nan | nan | nan | -0.396403 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.125583 | -0.369397 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.092181 | nan | nan | -1.111521 | nan | nan | nan | nan | nan | -0.339356 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.841087 | nan | nan | -1.239822 | -1.249646 | -0.604720 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -1.838380 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.247118 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.191007 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.669702 | nan | -0.626241 | nan | nan | nan | nan | nan | nan | -1.722196 | nan | -0.429002 | nan | nan | -0.454920 | nan | 0.217270 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | AASKLGVPQPAQRDQVNCQLYAVQPNDNCIDISSKNNITYAQLLSWNPSLSSTCSNLASLNSSSICVSNPKGTFSISSNTVGATDIATTTAPVPSPTLDQTTSRCAKYYQVSDGDDCSHLTAQFAITLKDFIFLNSEVWQNCTNLELGYYYCVEPVGYISTYPGYLPTATTKPFNQTSATSLPYDGDPWARFSSNSSVIPIANGTRVDCYSYVYVKNLTENLFADCWNMASMYEITREELVLWNPSLGNDGSSGSNLSGAEASQVASSIAIPTTAPSSITTNLYTYPCTVAANISYCVALVSSTGALPTNTAPPGPHASGEISNCTAWFAPEAYDTCKSILDIFEMSFANFYKMNPSVGPDCSGLAVGTNYCVSTYPNGEDPNDDWDGDDSIPSISPTGIITPTPTQSGMVSNCNKFYDVHSNDGCSAIASSQHVDLSSLYKWNPAIKTDCSGLQASVYVCIGILTTSMSTTSKLPTTTSKPSTTSKPPTGITTPTPTQSGMVKNCNKFYDVHSGDGCSAIASSQKVNLSSFYLWNPAVKTDCSGLQASVFVCIGTLTTSMSTTTSKLPTTTSKPSTTSKPPTGIITPTPTQNGMVKNCNKFYDVHSGDGCSAIASSQKVNLSSFYLWNPAVKTDCSGLQASVFVCVGTATTTTAAGITTPTPTQSGMVSGCNKFYDVHAGDGCSAIASSQKIALSSLYKWNPAVKTDCSGLQASVYICIGVGNAAAARVTGAAASFLEQKLISEEDLNSAVDHHHHHH | TAL6-6LysM |
| 4 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.352142 | 0.244700 | nan | -0.461016 | -0.234837 | -0.325556 | -0.052849 | nan | nan | -0.291694 | nan | -0.201436 | 0.068008 | 0.284881 | nan | -0.207676 | nan | nan | nan | nan | nan | nan | -0.509240 | 0.261744 | nan | nan | nan | -0.461065 | nan | 0.281879 | 0.314454 | nan | -0.547046 | nan | 0.065653 | nan | -0.206745 | nan | nan | nan | nan | nan | 0.069357 | -0.440782 | nan | nan | nan | -0.352217 | -0.313786 | -0.054417 | 0.220397 | 0.000593 | -0.468453 | nan | -0.642583 | nan | nan | nan | nan | nan | -0.213253 | 0.004967 | nan | -0.418988 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.019801 | nan | nan | nan | 0.348693 | -0.167535 | nan | nan | nan | nan | nan | nan | -0.072314 | nan | nan | nan | -0.415624 | nan | nan | nan | nan | -0.007247 | -0.096734 | 0.493524 | nan | -0.049574 | nan | nan | nan | nan | nan | nan | 0.356735 | -0.231039 | nan | nan | nan | 0.687253 | nan | nan | nan | nan | nan | nan | 0.039245 | nan | nan | nan | nan | 0.289550 | -0.017010 | 0.552749 | 0.253952 | 0.024845 | nan | nan | -0.092765 | nan | -0.050599 | nan | -0.253661 | nan | -0.034002 | -0.640639 | -0.332807 | nan | nan | -0.456518 | 0.013717 | nan | -0.257121 | -0.423334 | 0.238793 | 1.040417 | -0.252777 | nan | nan | nan | nan | -0.459521 | nan | 8.334515 | -0.000602 | 0.304265 | -0.082544 | -0.571446 | nan | 0.359063 | nan | nan | nan | -0.372296 | nan | nan | nan | -0.115188 | nan | -0.467803 | nan | -0.495880 | -0.284533 | -0.135805 | nan | nan | nan | 0.521431 | -0.180462 | nan | nan | -0.311932 | -0.437642 | -0.499129 | -0.580729 | -0.433832 | nan | nan | nan | -0.622161 | nan | 0.178139 | nan | -0.593969 | -0.479843 | -0.404232 | nan | nan | 0.426017 | 0.186881 | nan | -0.169694 | -0.266457 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.062682 | nan | nan | nan | nan | nan | nan | nan | nan | -0.087707 | -0.428183 | 0.186681 | nan | -0.260413 | -0.097686 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.084458 | nan | nan | nan | nan | nan | nan | nan | -0.236920 | nan | nan | nan | 0.000047 | nan | nan | nan | nan | -0.328665 | nan | nan | nan | nan | nan | nan | nan | nan | -0.050001 | nan | nan | nan | nan | nan | 0.096289 | -0.713843 | 0.626579 | 0.216825 | nan | nan | 0.025860 | nan | nan | nan | 0.140275 | nan | nan | nan | nan | nan | -0.483201 | -0.125274 | nan | nan | nan | 19.126138 | nan | nan | nan | nan | 0.447124 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.091157 | nan | nan | 0.236213 | nan | 0.958505 | -0.137814 | 0.299341 | -0.456594 | -0.101001 | 0.394345 | nan | -0.381016 | 0.230053 | -0.413905 | nan | 0.708616 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.731750 | nan | nan | nan | nan | nan | 0.055039 | nan | nan | nan | -0.425287 | -0.560030 | nan | nan | nan | -0.214257 | nan | nan | nan | nan | nan | 1.285163 | nan | nan | nan | nan | nan | -0.247194 | nan | nan | 0.738753 | 0.260701 | 0.550376 | nan | -0.507198 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.095765 | nan | 0.431800 | nan | nan | nan | nan | nan | nan | 0.055439 | nan | nan | nan | nan | nan | nan | 0.251386 | nan | nan | nan | nan | -0.593213 | nan | nan | -0.057296 | nan | -0.520796 | -0.694564 | nan | -0.697927 | nan | -0.489050 | nan | nan | nan | nan | -0.148911 | nan | nan | nan | nan | -0.500282 | nan | nan | nan | nan | nan | nan | nan | 0.060701 | nan | 0.216923 | nan | nan | nan | -0.414411 | 2.763139 | 0.261430 | -0.152436 | -0.152840 | -0.324842 | nan | nan | nan | nan | nan | 0.024281 | nan | -0.436718 | 0.029816 | 0.086215 | 0.558052 | nan | nan | 0.752209 | nan | nan | nan | nan | 0.016113 | nan | nan | -0.574978 | nan | nan | nan | 0.183228 | nan | 0.028883 | -0.388068 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.558211 | 0.161656 | -0.273586 | nan | -0.287044 | nan | nan | nan | nan | nan | nan | nan | -0.203313 | 0.303259 | nan | 0.610638 | nan | -0.080314 | nan | nan | nan | -0.228164 | nan | nan | nan | nan | nan | 0.188941 | nan | nan | nan | -0.316051 | nan | -0.200855 | nan | nan | nan | nan | nan | nan | nan | nan | 0.765834 | nan | -0.060499 | nan | nan | -0.338792 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.474869 | -0.312395 | -0.450562 | nan | nan | 0.631150 | -0.184554 | nan | -0.496568 | nan | 1.025249 | -0.191439 | nan | nan | -0.393800 | nan | -0.014128 | -0.392889 | -0.182935 | -0.322916 | nan | 0.109831 | -0.253569 | -0.304187 | -0.460293 | nan | nan | nan | nan | -0.153168 | nan | nan | nan | nan | -0.545048 | -0.093180 | nan | nan | nan | nan | nan | nan | 0.225497 | nan | nan | nan | nan | nan | nan | nan | -0.050547 | nan | -0.429489 | -0.035623 | nan | -0.651230 | -0.757136 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.407953 | -0.172880 | 0.225896 | -0.124650 | 0.404438 | -0.278338 | nan | 0.676358 | 0.102604 | nan | nan | nan | 0.071175 | nan | nan | nan | nan | 0.701722 | 2.766895 | -0.332718 | nan | nan | nan | nan | nan | nan | 0.200391 | -0.113778 | 0.090591 | nan | nan | nan | -0.342221 | 0.440389 | 0.968416 | 0.152312 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.438715 | nan | nan | 0.116453 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.289726 | nan | nan | nan | nan | nan | nan | nan | -0.127844 | nan | nan | nan | nan | nan | nan | 0.135441 | -0.732415 | nan | nan | nan | nan | 0.369340 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.234051 | nan | -0.376654 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.426711 | nan | nan | 0.329170 | nan | 0.088458 | nan | nan | nan | nan | nan | 0.203284 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.156843 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.970065 | nan | nan | nan | nan | nan | nan | -0.098160 | nan | nan | 0.014632 | nan | nan | nan | nan | 0.256873 | -0.049152 | -0.260200 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.201269 | -0.342285 | 0.110783 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.158687 | nan | -0.453498 | nan | nan | nan | nan | nan | -0.354170 | nan | nan | nan | -0.111170 | -0.062455 | nan | nan | nan | 0.170343 | 0.024254 | -0.335108 | nan | nan | nan | nan | 0.156729 | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.557843 | nan | nan | nan | nan | nan | 1.352004 | nan | nan | nan | -0.020119 | -0.209953 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.418717 | nan | nan | 0.107102 | -0.168479 | -0.042227 | nan | nan | 0.046235 | nan | 0.253850 | nan | nan | nan | nan | nan | -0.084545 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 1.178867 | -0.103371 | nan | 1.540236 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.000352 | nan | nan | nan | -0.085181 | nan | nan | -0.485347 | nan | nan | nan | nan | nan | nan | nan | nan | -0.021827 | nan | -0.113681 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.366496 | nan | nan | nan | nan | nan | nan | -0.367804 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.119126 | nan | -0.416085 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.005896 | 0.998665 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.217622 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.041767 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.163504 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.486207 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.865622 | nan | nan | nan | nan | nan | nan | 0.245497 | nan | nan | nan | nan | nan | nan | nan | -0.573961 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.266594 | nan | nan | nan | nan | nan | nan | nan | -0.025309 | nan | 0.024376 | -0.275329 | 0.184680 | -0.187755 | -0.270358 | 0.595347 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.121682 | nan | nan | 0.906455 | nan | nan | nan | nan | nan | nan | -0.386199 | nan | nan | 0.166275 | -0.364857 | -0.473669 | nan | nan | nan | nan | -0.298895 | -0.150466 | 0.157226 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.082997 | nan | 1.106571 | nan | nan | nan | nan | nan | -0.391481 | -0.357996 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.370765 | nan | -0.639713 | -0.205941 | nan | 0.170073 | -0.233150 | -0.005755 | 0.082972 | 0.155363 | nan | 0.054915 | nan | nan | nan | 0.130737 | 0.848404 | -0.141295 | nan | nan | nan | -0.146291 | -0.317983 | 0.005869 | nan | -0.225642 | -0.027208 | 0.133381 | nan | -0.503229 | nan | nan | nan | 0.500372 | nan | nan | 0.147946 | 0.223088 | -0.227036 | -0.085662 | nan | nan | nan | nan | nan | nan | -0.484465 | nan | -0.187568 | nan | nan | nan | -0.192372 | nan | nan | -0.304203 | nan | -0.543672 | nan | nan | nan | nan | nan | nan | nan | -0.192604 | nan | nan | -0.421335 | -0.575204 | nan | -0.451805 | 0.044478 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.018559 | 0.154486 | nan | 0.491860 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.262406 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.433638 | nan | nan | nan | -0.316137 | nan | -0.005388 | -0.184509 | nan | nan | -0.263857 | nan | nan | nan | nan | nan | nan | nan | -0.417204 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.467643 | nan | nan | nan | nan | 0.464403 | -0.190363 | nan | nan | nan | nan | nan | -0.562790 | -0.086740 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.281332 | nan | nan | nan | nan | 0.048001 | nan | nan | nan | nan | nan | nan | 0.074367 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.003137 | nan | nan | nan | nan | nan | nan | nan | nan | -0.204196 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.441238 | nan | nan | nan | nan | nan | nan | nan | nan | 0.017722 | -0.617849 | nan | nan | nan | -0.053200 | nan | nan | -0.355265 | nan | nan | nan | nan | nan | 0.305782 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.382800 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.316541 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.313869 | nan | nan | nan | 1.938720 | nan | nan | -0.170564 | nan | nan | nan | -0.365692 | -0.259854 | -0.271620 | nan | nan | nan | nan | -0.033167 | -0.425726 | nan | -0.508203 | -0.337518 | nan | nan | nan | nan | -0.321046 | nan | 0.109863 | nan | -0.085295 | nan | nan | -0.392853 | -0.332886 | 0.046866 | nan | nan | -0.072946 | -0.282880 | nan | 0.437903 | nan | -0.390503 | -0.237883 | -0.445866 | nan | nan | nan | nan | nan | nan | nan | -0.697174 | nan | nan | nan | 0.996852 | -0.021289 | -0.249012 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.551922 | -0.039140 | -0.235864 | 0.847094 | nan | -0.440410 | nan | nan | nan | 0.132761 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.000585 | -0.388119 | nan | nan | nan | nan | nan | nan | -0.267915 | nan | 0.117292 | nan | nan | nan | nan | -0.729473 | nan | nan | 0.703680 | nan | nan | nan | nan | nan | -0.258475 | nan | nan | nan | nan | nan | nan | nan | 0.516386 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.510212 | -0.103087 | -0.213671 | 0.411249 | nan | nan | nan | 0.717649 | nan | -0.228116 | nan | nan | nan | -0.192381 | nan | nan | nan | -0.224747 | nan | nan | -0.543443 | nan | -0.405085 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.246832 | 0.056778 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.134523 | nan | nan | 0.266096 | nan | nan | nan | -0.053077 | nan | nan | 1.759381 | nan | nan | nan | nan | nan | nan | 0.512064 | nan | nan | 0.537340 | -0.150449 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.363980 | nan | nan | nan | -0.363345 | nan | nan | 0.234002 | -0.166385 | nan | nan | nan | nan | nan | nan | nan | -0.539494 | 0.103373 | nan | nan | nan | nan | nan | nan | nan | -0.481606 | nan | nan | nan | nan | nan | nan | 0.465333 | 0.099508 | nan | nan | nan | nan | nan | nan | nan | -0.502997 | nan | nan | nan | nan | nan | nan | nan | -0.225889 | 0.493369 | -0.622523 | -0.385000 | nan | nan | -0.153559 | nan | nan | -0.134633 | nan | nan | nan | nan | -0.362093 | -0.228914 | nan | nan | nan | -0.264698 | 0.105541 | -0.079860 | nan | nan | -0.263121 | nan | -0.067154 | nan | nan | nan | -0.140849 | nan | -0.498545 | nan | 0.202228 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.739133 | -0.509332 | nan | nan | nan | 0.258930 | nan | -0.679921 | nan | 0.228373 | nan | nan | nan | nan | nan | nan | nan | -0.319420 | nan | nan | nan | nan | nan | nan | -0.157416 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.371906 | -0.294867 | nan | nan | nan | -0.195848 | nan | nan | nan | nan | 0.107341 | nan | nan | nan | -0.619119 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.179686 | nan | nan | -0.236770 | nan | -0.564975 | nan | -0.423714 | nan | nan | nan | nan | nan | -0.194092 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.359516 | -0.736095 | -0.451625 | nan | nan | nan | -0.508315 | -0.452826 | nan | nan | -0.247064 | nan | nan | nan | -0.496840 | 0.029668 | nan | nan | nan | nan | nan | nan | -0.384303 | -0.345193 | nan | nan | -0.082610 | nan | nan | nan | nan | -0.466420 | nan | nan | nan | nan | nan | -0.123249 | -0.508771 | -0.048365 | nan | nan | nan | nan | -0.300055 | nan | nan | nan | nan | nan | nan | nan | -0.365758 | nan | nan | nan | nan | nan | nan | nan | nan | -0.209880 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.305390 | -0.462766 | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.509763 | nan | nan | -0.390991 | nan | nan | nan | nan | nan | -0.311137 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.105759 | nan | nan | -0.411670 | -0.099623 | -0.187737 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.181018 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | 0.629400 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.371103 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | -0.285324 | nan | 0.013751 | nan | nan | nan | nan | nan | nan | -0.427397 | nan | -0.077225 | nan | nan | -0.087638 | nan | 0.645254 | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | nan | ACNNEWEDEQYEQYISFKSPIPAGGEGVTDIYVRYKEDGKVTYRLPIVVSGSTTNTLDREVHIAVDKDTLETLNIERFSIHRPGLWYTALEEDKYDFPKTVHISAGSSVEQLNIDFNLQGIDMLEKWVLPLTIVDDPSYNYKSHPRKNYAKALLRVIPFNNYSGSYTSSAMKVYTYINGKPDTNARTANKRTGYVINDNTVFFYAGLISEDLDKEVREKYKINVHFNEDGTLHMEQGDPTNKMKFELIGTPTYSKTSVMDATRPYLERRYVQIMFEYDFEDFSYGGNGVDVIPIKYKVEGSMTLQRNINTQIPDEDQQIEW | SP15308A-bot-339-19-339 |
As you can see, this dataset contains numerical values (Z-scores) corresponding to the ability of different lectins to bind different glycans. From these data, we can compute a heatmap with get_heatmap. The results should teach us whether some specific strains bind specific glycans. As we work with quantitative values, the datatype parameter must be set to 'response'. If you want to include literature-annotated motifs (e.g., VIM, Lewis X, Blood group A), just add ‘known’ to the list of the feature_set. Let’s see what kind of heatmap we can generate using this dataset.
get_heatmap(glycan_binding.iloc[:,:-2], motifs = True, feature_set = ['exhaustive'], datatype = 'response', yticklabels = 1,
xticklabels = False)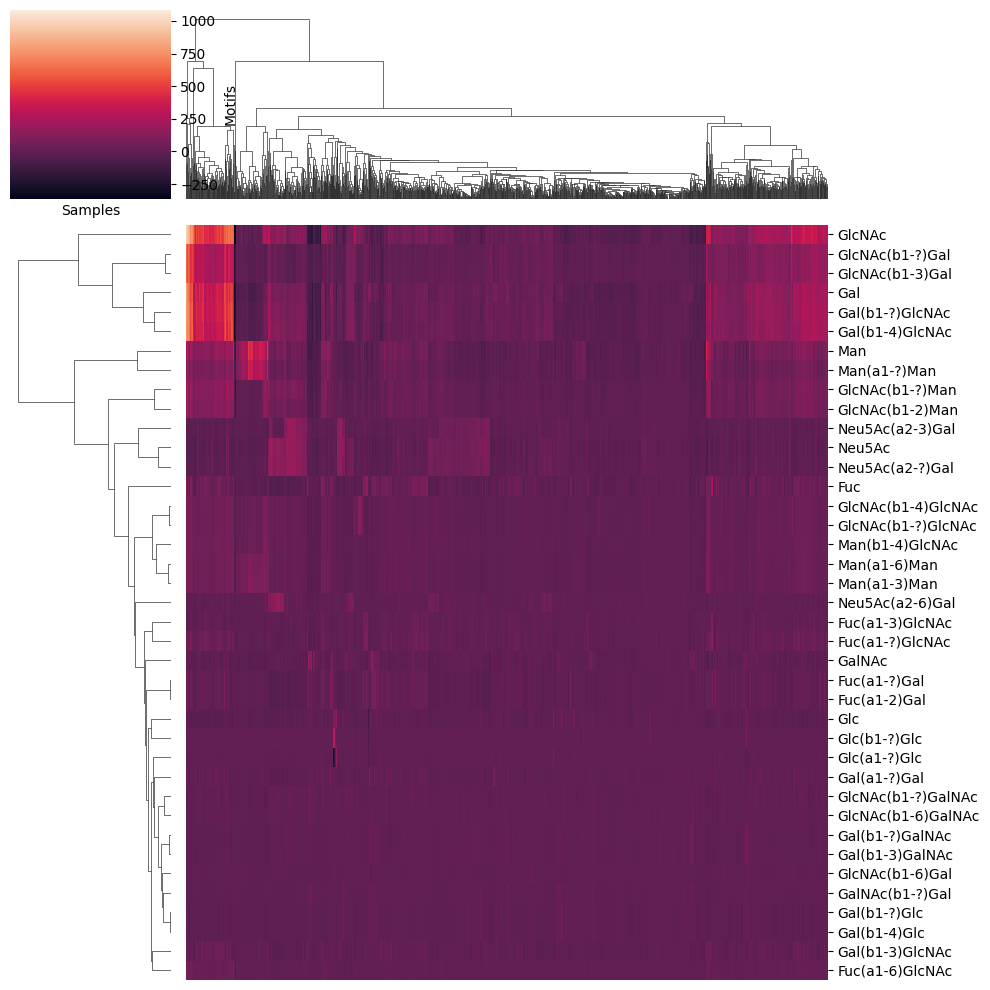
Due to the high number of lectins in this dataset, the heatmap is difficult to analyze in detail. However, we can clearly see that there are clusters of lectins recognizing specific glycan motifs. Examples include Neu5Ac(a2-6)Gal, Neu5Ac(a2-3)Gal, Fuc(a1-6)GlcNAc, and others. All these represent well-known glycan motifs with important roles in biology.
Another way to analyze the glycan binding capacities of lectins is to use glycowork.motif.analysis.get_pvals_motifs. As we are working with data obtained from different proteins, we must also set multiple_samples to True.
results = get_pvals_motifs(glycan_binding.iloc[:,:-2].T.reset_index(), feature_set = ['exhaustive'], multiple_samples = True)| motif | pval | corr_pval | effect_size | |
|---|---|---|---|---|
| 504 | Gal(b1-4)GlcNAc | 0.0 | 0.0 | 0.19871392612060212 |
| 740 | Neu5Ac(?2-2/3/6)Gal | 0.0 | 0.0 | 0.1940171392764387 |
| 133 | GlcNAc | 0.0 | 0.0 | 0.18544051329412226 |
| 410 | Gal(?1-3/4/6)GlcNAc | 0.0 | 0.0 | 0.1781652222810397 |
| 199 | Man | 0.0 | 0.0 | 0.17669180124256265 |
| 522 | Man(a1-3)Man | 0.0 | 0.0 | 0.16997960952416624 |
| 753 | Sia(?2-2/3/6)Gal | 0.0 | 0.0 | 0.16976553648570486 |
| 330 | Man(?1-2/3/4/6)Man | 0.0 | 0.0 | 0.1694073251729428 |
| 1202 | Man(a1-6)Man | 0.0 | 0.0 | 0.16651100439280453 |
| 1109 | GlcNAc(b1-3)Gal | 0.0 | 0.0 | 0.15769567508257473 |
| 225 | Neu5Ac | 0.0 | 0.0 | 0.15640433487463892 |
| 46 | Gal | 0.0 | 0.0 | 0.15384314048457948 |
| 665 | Man(b1-4)GlcNAc | 0.0 | 0.0 | 0.15319917505367894 |
| 1097 | Man(?1-3/4)GlcNAc | 0.0 | 0.0 | 0.15305859223388624 |
| 291 | GlcNAc(?1-2/3/4/6)Gal | 0.0 | 0.0 | 0.15173728781342968 |
| 272 | Sia | 0.0 | 0.0 | 0.14123800007994433 |
| 457 | GlcNAc(b1-4)GlcNAc | 0.0 | 0.0 | 0.1398511509027576 |
| 329 | Neu5Ac(a2-3)Gal | 0.0 | 0.0 | 0.13730243672190867 |
| 685 | GlcNAc(b1-2)Man | 0.0 | 0.0 | 0.13714727818087413 |
| 391 | GlcNAc(?1-3/4/6)GlcNAc | 0.0 | 0.0 | 0.13694071609068692 |
| 519 | GlcNAc(?1-2/3/4/6)Man | 0.0 | 0.0 | 0.13419627571413126 |
| 1031 | Neu5Ac(a2-6)Gal | 4.1753444778352537e-308 | 1.5908062460552316e-306 | 0.1287068602531006 |
| 1153 | Sia(a2-6)Gal | 0.0 | 0.0 | 0.12250056603524426 |
| 776 | Sia(a2-3)Gal | 0.0 | 0.0 | 0.11422628882983418 |
| 1266 | GlcNAc(b1-6)Man | 1.6619951525863255e-255 | 5.5872366453122646e-254 | 0.09228052805408492 |
These results indicate, for each motif, a p-value and a corrected p-value. These quantitative results support our qualitative observations from before. However, this represents a bird’s eye view across all lectins in the dataset. A follow-up analysis might be to focus on a specific group of lectins, analyzing their binding patterns in depth.
Example 3: Glycan classification using machine learning models
The investigation of glycans from animals and viruses demonstrated how specific their presence in an organism can be. This particular characteristic can be exploited to predict the source of a given glycan among organisms. By classifying which glycans stem from which species/taxonomic groups, machine learning models can be trained to learn which motifs are enriched in which species, allowing for further insights.
Let’s see how we can use glycowork to classify human versus non-human primate glycans! First, we need to annotate each known glycan using a binary code where 1 means human glycan and 0 means non-human primate glycan.
human = [1 if k == 'Homo_sapiens' else 0 for k in df_species[df_species.Order=='Primates'].Species.values.tolist()]Now, we have to generate two sets of glycans that we will use as train set and test set. In addition, we also need their corresponding sets with classes labelled as 0 or 1. One way to do it is to use glycowork.ml.train_test_split.general_split. This function assigns the data with a 80/20 ratio to train and test sets respectively.
X_train, X_test, y_train, y_test = general_split(df_species[df_species.Order=='Primates'].glycan.values.tolist(), human)It is now time to train a machine learning model on these data! Different machine learning algorithms are available in glycowork and here we can use glycowork.ml.model_training.train_ml_model. This classification method uses a standard machine learning model (XGBoost) trained on short glycan motifs which may not be the most accurate, but can be sufficient for a preliminary exploration of our problem. Plus, it has the advantage of running faster than other more complex algorithms such as deep learning so… let’s give it a try!
model_ft, _, X_test = train_ml_model(X_train, X_test, y_train, y_test, feature_calc = True, feature_set = ['terminal'],
return_features = True)
Calculating Glycan Features...
Training model...
Evaluating model...
Accuracy of trained model on separate validation set: 0.8570190641247833As you can see, even this (quite) simple algorithm is enough to reach high prediction accuracy on the test set! It also confirms that glycan sequence is highly representative of their origin and allows to efficiently discriminate human from non-human primate glycans.
One more step we can do is the analysis of the model we have generated. This is important, as we do not know yet what information is the most useful for the model to predict the origin of a glycan. The glycowork.ml.model_training.analyze_ml_model function can help us to understand on which basis the model predictions are made.
Interesting! It appears that the presence/absence of alpha-Gal and Neu5Gc are quite discriminative of its belonging to Homo sapiens or not!
Next to knowing what is important for model prediction, it is often also crucial to know the limits of a model. In glycowork, you can use glycowork.ml.model_training.get_mismatch module to get a sense of wrong classifications. This could for instance inform you about potential model biases.
get_mismatch(model_ft, X_test, y_test)[('Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc-ol', 0.666021466255188),
('Gal(b1-4)GlcNAc(b1-2)Man(a1-6)[GlcNAc(b1-2)Man(a1-3)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
0.8374393582344055),
('Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-3)Gal(b1-4)Glc-ol', 0.8031653165817261),
('Neu5Ac(a2-3)Gal(b1-3/4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
0.9542927145957947),
('Gal(b1-3)[Neu5Ac(a2-6)]GlcNAc(b1-3)Gal(b1-4)Glc-ol', 0.7572895884513855),
('Fuc(a1-3/4)[Gal(b1-3/4)]GlcNAc(b1-2)[Fuc(a1-3/4)[Gal(b1-3/4)]GlcNAc(b1-4)]Man(a1-3)[Fuc(a1-3/4)[Gal(b1-3/4)]GlcNAc(b1-2)[Fuc(a1-3/4)[Gal(b1-3/4)]GlcNAc(b1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
0.7792727947235107),
('Gal(b1-3/4)GlcNAc(b1-2/4)[GlcNAc(b1-2/4)]Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
0.889919102191925),
('{Gal(b1-3/4)GlcNAc(b1-3)}{Gal(b1-3/4)GlcNAc(b1-3)}{Gal(b1-3/4)GlcNAc(b1-3)}{Gal(b1-3/4)GlcNAc(b1-3)}{Fuc(a1-2/3/4)}Fuc(a1-3)[Gal(b1-4)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-4)GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
0.3473208248615265),
('Gal(b1-4)Glc-ol', 0.6590997576713562),
('Gal(b1-4)GlcNAc(b1-2)Man(a1-3)[Man(a1-3)[Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
0.8307253122329712)]Here we can see that our model was especially confused by glycans which can occur in both types of orders and therefore are ambiguous for classification ambiguous. Maybe in these scenarios a more powerful, deep learning-based model might be able to differentiate (at least a bit better)!
Example 4: Constructing and exploring biosynthetic networks
The network module of glycowork is designed to readily construct and analyze biosynthetic networks from a list of glycans in a very modular way. Below, this is illustrated with a fictional set of milk glycans (all default parameters of construct_network are optimized for milk glycans and should be changed for other glycan classes). Just inputting a list of biosynthetically related glycans into glycowork.network.biosynthesis.construct_network and then glycowork.network.biosynthesis.plot_network will build and plot our biosynthetic network! (depending on your browser etc, this might not render correctly)
glycans = ["Gal(b1-4)Glc-ol", "GlcNAc(b1-3)Gal(b1-4)Glc-ol",
"GlcNAc6S(b1-3)Gal(b1-4)Glc-ol",
"Gal(b1-4)GlcNAc(b1-3)Gal(b1-4)Glc-ol", "Fuc(a1-2)Gal(b1-4)Glc-ol",
"Gal(b1-4)GlcNAc(b1-3)[Gal(b1-3)GlcNAc(b1-6)]Gal(b1-4)Glc-ol"]
network = construct_network(glycans)
plot_network(network)We can see that, in these networks, every dot is a glycan (and can be hovered over to see its sequence) and every link is the addition of a monosaccharide in a defined linkage by an enzyme. Blue dots are glycans which we provided as input, orange glycans are inferred in order to connect the network.
One thing you can immediately see is that there often are multiple paths to reach a blue dot, via several orange routes. This is because we don’t know the order in which the monosaccharides have been added. We show in recent work that this order is highly conserved. Therefore, we could use knowledge gathered over decades in the breast milk of over a hundred mammalian species, to find out which order is realistic.
glycowork can help you here as well, since it contains pre-computed biosynthetic networks for milk glycans of >100 mammalian species. We can use the glycowork.network.biosynthesis.evoprune_network function to use this information to prune our networks, removing paths that are evolutionarily very unlikely.
print(len(network.edges()))
network = evoprune_network(network)
print(len(network.edges()))
plot_network(network)14
7Nice, that’s a bit more organized! evoprune_network has removed a few of the nodes that had zero probability. You can see that the orange nodes have changed size as well. This is to indicate their probability. If you change the threshold parameter of evoprune_network these low-probability nodes might also be pruned away, yet network connectivity is always preserved.
Now, let’s say we want to highlight parts of the network containing the monosaccharide Fuc. For this, we can use glycowork.network.biosynthesis.highlight_network. With this function, you can highlight many things in a network: motifs, abundances, species occurence, evolutionary conservation, etc.
network = highlight_network(network, 'motif', motif = 'Fuc')
plot_network(network)You can see that the node color has changed now. Every glycan/node that contains Fuc now is showing up as a green node, while those lacking Fuc are purple. You can also still make out which nodes are inferred (transparent) and actually observed (solid). The probabilities from evoprune_networkare also still contained within the network. Apart from these visualizations, you can use these networks for all kinds of analytical work, from glycowork.motif.graph.generate_graph_features to glycowork.network.evolution.get_communities; your only limitation is your creativity!:-)
We can also use the functions within glycowork.network to compare species and their milk glycomes: Let’s have a look at the two pre-computed biosynthetic networks (found in net_dic) of black bears (Ursus americanus) and brown bears (Ursus arctos).
blackbear_network = net_dic['Ursus_americanus']
brownbear_network = net_dic['Ursus_arctos']
plot_network(network = brownbear_network)As in our example we have two networks, it is possible to directly combine and compare them to identify conserved and specific nodes, using for instance glycowork.network.biosynthesis.network_alignment.
combined_network = network_alignment(network_a = blackbear_network, network_b = brownbear_network)
plot_network(network = combined_network)In this new representation, the blue nodes are specific to network_a (here the black bear), and orange nodes are specific to network_b (the brown bear). The remaining brown nodes are conserved in both species. Virtual nodes follow the same color scheme but are transparent. Here, we can see many blue nodes that are specific to the black bear. This is not so surprising, as the black bear network is larger.
We can also make use of the network.evolution module to compare the similarity of related species, such as different kinds of bears. Using the metric of Jaccard distance between their networks, we can retrieve sequences and networks for various bear species within glycowork with the function glycowork.network.evolution.distance_from_metric. Then, we can use the distance matrix in the glycowork.network.evolution.dendrogram_from_distance function to plot the resulting clustering. One immediate result here is that polar bears (Ursus maritimus) seem to have a very different milk glycome from other types of bears!
bears = [net_dic[k] for k in set(list(net_dic.keys())) if 'Ursus' in k]
df_bears = df_species[df_species.Genus=='Ursus'].reset_index(drop = True)
dm = distance_from_metric(df_bears, bears, cut_off = 5)
dendrogram_from_distance(dm)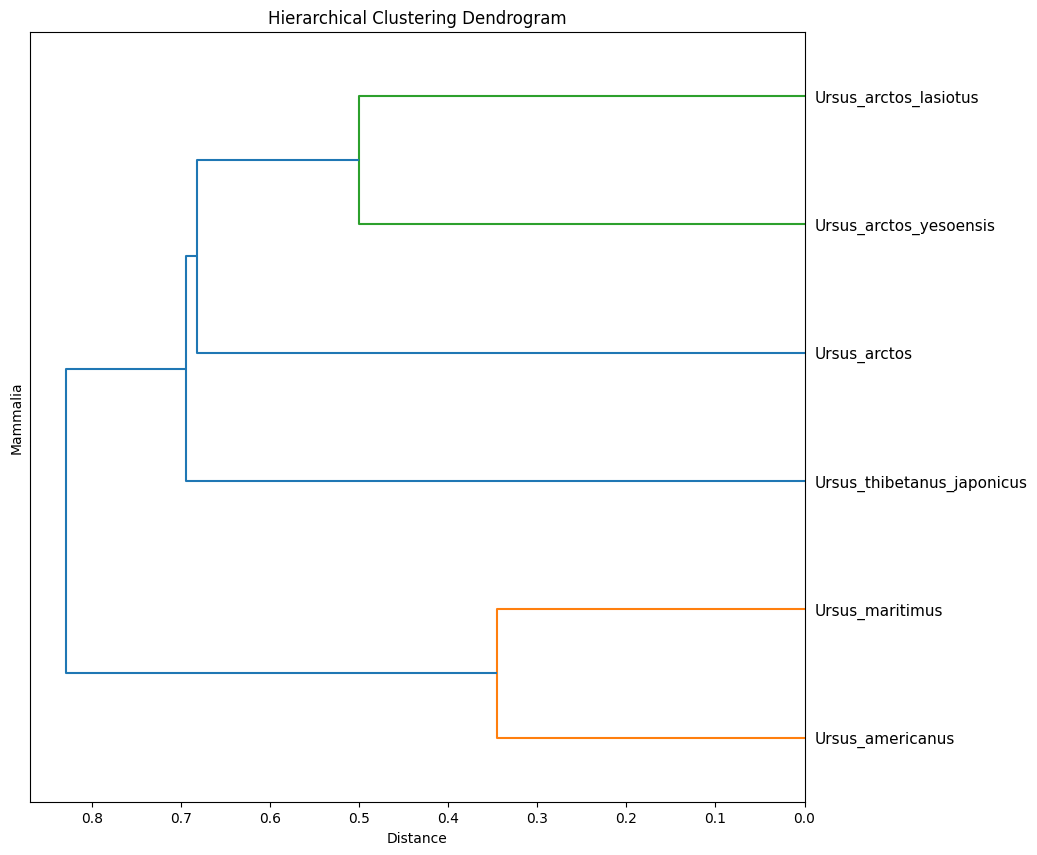
| Ursus_americanus | Ursus_thibetanus_japonicus | Ursus_maritimus | Ursus_arctos | Ursus_arctos_yesoensis | Ursus_arctos_lasiotus | |
|---|---|---|---|---|---|---|
| Ursus_americanus | 0.000000 | 0.681818 | 0.500000 | 0.854167 | 0.867470 | 0.828571 |
| Ursus_thibetanus_japonicus | 0.681818 | 0.000000 | 0.721311 | 0.693878 | 0.829545 | 0.852174 |
| Ursus_maritimus | 0.500000 | 0.721311 | 0.000000 | 0.850000 | 0.866667 | 0.882353 |
| Ursus_arctos | 0.854167 | 0.693878 | 0.850000 | 0.000000 | 0.940299 | 0.958333 |
| Ursus_arctos_yesoensis | 0.867470 | 0.829545 | 0.866667 | 0.940299 | 0.000000 | 0.345238 |
| Ursus_arctos_lasiotus | 0.828571 | 0.852174 | 0.882353 | 0.958333 | 0.345238 | 0.000000 |
Example 5: Differential glycomics analyses
Let’s say you have collected glycomics data from healthy and diseased samples and now you want to find out what’s really different between the two. There are many ways that you can go about this, depending on your exact question but the most likely answer to your question can be found in the glycowork.motif.analysis.get_differential_expression function. This function allows you to test whether any sequences (or motifs) are exhibiting significantly different abundances between your conditions.
But first, we need some data! Now, maybe you have your own data here but you can also easily access the glycomics data stored within glycowork via the glycowork.glycan_data.loader.glycomics_data_loader. Using dir(glycomics_data_loader) will tell you which datasets are currently available (hint: there are also analogous dataloaders for glycoproteomics and lectin microarray data). This example will work with a skin cancer O-glycomics dataset but feel free to use your own data if you have it.
df = glycomics_data_loader.human_skin_O_PMC5871710| glycan | control_1 | tumor_1 | control_2 | tumor_2 | control_3 | tumor_3 | control_4 | tumor_4 | control_5 | tumor_5 | control_6 | tumor_6 | control_7 | tumor_7 | control_8 | tumor_8 | control_9 | tumor_9 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | GalOS(b1-3)[Neu5Ac(a2-6)]GalNAc | 2.755913 | 0.620301 | 3.019806 | 4.148615 | 2.413021 | 0.424605 | 1.764977 | 0.744528 | 4.518547 | 0.849240 | 3.174455 | 1.543922 | 3.752813 | 1.287601 | 1.574444 | 0.627539 | 4.415230 | 0.425039 |
| 1 | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 0.775234 | 0.178490 | 0.778083 | 0.832907 | 0.824018 | 0.410873 | 0.617573 | 2.133773 | 0.603573 | 0.544599 | 1.317444 | 1.256997 | 0.860332 | 0.340373 | 0.927956 | 0.494144 | 1.045950 | 0.590404 |
| 2 | Neu5Ac(a2-3)Gal(b1-3)GalNAc | 9.219494 | 8.436141 | 16.190721 | 11.032313 | 16.061363 | 8.133529 | 12.291188 | 10.143781 | 10.865172 | 14.353894 | 12.313024 | 11.601938 | 13.246380 | 8.352032 | 12.352587 | 9.731834 | 16.170263 | 12.693331 |
| 3 | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 2.770825 | 0.944913 | 1.531698 | 2.427562 | 1.570204 | 2.066165 | 2.832475 | 2.508954 | 2.228299 | 1.757840 | 4.811796 | 3.641077 | 1.437487 | 1.470697 | 2.324956 | 2.192553 | 0.805130 | 1.100490 |
| 4 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 35.912676 | 30.058418 | 48.420833 | 43.738436 | 46.692847 | 26.728713 | 39.971128 | 32.290024 | 51.472292 | 40.669153 | 41.927351 | 37.755641 | 43.701528 | 32.565498 | 47.145757 | 38.565763 | 44.683562 | 33.532942 |
| 5 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-8)Neu5Ac(a2-6)]GalNAc | 0.664610 | 0.495280 | 1.451470 | 1.289237 | 1.331164 | 0.441114 | 0.657772 | 0.330536 | 0.654086 | 0.219399 | 1.433102 | 0.791533 | 0.722264 | 0.589715 | 1.337691 | 0.779090 | 1.734299 | 0.605036 |
| 6 | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 1.069934 | 1.735744 | 1.468105 | 1.509499 | 1.128653 | 1.812381 | 0.816789 | 6.345353 | 2.179460 | 1.261105 | 1.438459 | 4.410718 | 0.669910 | 1.310332 | 1.838577 | 0.989791 | 1.329698 | 1.392818 |
| 7 | Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc6S(b1-6)]GalNAc | 0.599954 | 1.213936 | 0.450138 | 0.469515 | 0.528332 | 2.891749 | 0.770085 | 0.853955 | 0.853148 | 4.776427 | 0.669361 | 1.323619 | 0.914651 | 1.027152 | 1.633415 | 4.333793 | 0.365928 | 3.162330 |
| 8 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Gal(b1-3)]GalNAc | 6.005921 | 6.365952 | 5.623333 | 5.965819 | 5.299405 | 7.027606 | 10.523154 | 12.570391 | 4.753029 | 6.895145 | 12.325022 | 10.338465 | 5.401390 | 5.785955 | 7.054084 | 5.775716 | 11.549820 | 4.633692 |
| 9 | Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]GalNAc | 3.557548 | 2.179078 | 2.341708 | 2.594836 | 2.408925 | 1.838783 | 3.403741 | 1.561446 | 1.909720 | 0.838742 | 1.389057 | 1.590800 | 2.557034 | 3.846191 | 0.765387 | 0.660688 | 1.428603 | 0.845144 |
| 10 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 8.448576 | 14.890684 | 4.765329 | 7.229355 | 5.669282 | 18.752163 | 7.503251 | 7.395700 | 6.107556 | 8.367088 | 7.748875 | 8.248336 | 9.086158 | 13.802771 | 6.255983 | 9.265805 | 4.196432 | 11.075915 |
| 11 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)Gal(b1-3)]GalNAc | 25.222604 | 26.314089 | 12.026590 | 16.037174 | 13.649831 | 21.493736 | 16.473606 | 19.289899 | 10.427178 | 12.014370 | 9.666302 | 14.461389 | 15.276941 | 25.481534 | 12.897587 | 16.978625 | 10.428889 | 18.751042 |
| 12 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Neu5Ac(a2-3)Gal(b1-3)]GalNAc | 2.996712 | 6.566972 | 1.932188 | 2.724731 | 2.422956 | 7.978585 | 2.374260 | 3.831659 | 3.427941 | 7.452997 | 1.785750 | 3.035567 | 2.373113 | 4.140150 | 3.891578 | 9.604660 | 1.846197 | 11.191816 |
We can see that the dataset is structured as having the glycans in the very first column and then every sample being another column. This is the standard set-up that most functions in glycowork.motif.analysis expect to get. Note that, while glycans here are written in IUPAC-condensed, you can use pretty much any common glycan nomenclature (WURCS, GlycoCT, IUPAC-extended, etc.) and our internal conversions will make it work.
Now glycowork.motif.analysis.get_differential_expression basically only needs to know two things: (i) your dataset and (ii) which columns belong to which condition. There are many more options and settings that you can choose, but that’s the minimum. Our example dataset tell us in the column headings which sample comes from a tumor and which from a healthy setting. Our function now wants to know either the position of the column or the column names of both the control samples and the disease samples. We will further use one optional setting, “paired = True”, to indicate that these samples are matched (control_1 and tumor_1 come from the same patient). Analyzing paired samples in a paired manner is not only more sensitive but also more appropriate, statistically speaking.
So, let’s see which glycans are differentially abundant in skin cancers!
get_differential_expression(df, [1,3,5,7,9,11,13,15,17], [2,4,6,8,10,12,14,16,18], paired = True)You're working with an alpha of 0.05632533070968504 that has been adjusted for your sample size of 18.
Significance inflation detected. The CLR/ALR transformation possibly cannot handle this dataset. Consider running again with a higher gamma value. Proceed with caution; for now switching to Bonferroni correction to be conservative about this.| Glycan | Mean abundance | Log2FC | p-val | corr p-val | significant | corr Levene p-val | Effect size | Equivalence p-val | |
|---|---|---|---|---|---|---|---|---|---|
| 7 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 39.974308 | -3.694140 | 3.536872e-12 | 4.597933e-11 | True | 0.010221 | -21.631303 | 1.000000 |
| 1 | Gal(b1-3)[Neu5Ac(a2-6)]GalNAc | 0.775084 | 2.004187 | 5.691316e-08 | 7.398710e-07 | True | 0.005665 | 6.388785 | 1.000000 |
| 4 | Neu5Ac(a2-3)Gal(b1-3)GalNAc | 11.907405 | -1.945773 | 7.023338e-08 | 9.130339e-07 | True | 0.008918 | -6.219744 | 1.000000 |
| 10 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Neu5Ac(a2-3)... | 16.574094 | -1.975481 | 5.506862e-07 | 7.158920e-06 | True | 0.005665 | -4.774636 | 1.000000 |
| 6 | Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc6S(b1-6)]... | 1.478294 | 2.273060 | 1.008567e-06 | 1.311137e-05 | True | 0.006427 | 4.414143 | 1.000000 |
| 8 | Neu5Ac(a2-3)Gal(b1-3)[Neu5Ac(a2-8)Neu5Ac(a2-6)... | 0.855548 | 1.730360 | 1.314023e-05 | 1.708230e-04 | True | 0.001913 | 3.143080 | 1.000000 |
| 0 | Fuc(a1-2)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 1.731629 | 1.425387 | 1.609128e-05 | 2.091866e-04 | True | 0.006427 | 3.058077 | 1.000000 |
| 9 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]Ga... | 8.683615 | -0.893815 | 2.063204e-05 | 2.682165e-04 | True | 0.006427 | -2.956445 | 1.000000 |
| 11 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Gal(b1-3)]... | 7.469484 | -1.086662 | 3.133806e-04 | 4.073948e-03 | False | 0.008977 | -2.009422 | 1.000000 |
| 2 | Gal(b1-4)GlcNAc(b1-6)[Gal(b1-3)]GalNAc | 2.088085 | 0.732280 | 4.319286e-03 | 5.615072e-02 | False | 0.002963 | 1.312045 | 1.000000 |
| 12 | Neu5Ac(a2-3)Gal(b1-4)GlcNAc6S(b1-6)[Neu5Ac(a2-... | 4.361909 | 0.475682 | 8.921154e-03 | 1.159750e-01 | False | 0.004764 | 1.144185 | 1.000000 |
| 5 | Neu5Ac(a2-3)Gal(b1-3)[Gal(b1-4)GlcNAc(b1-6)]Ga... | 1.982425 | 0.780486 | 9.781398e-03 | 1.271582e-01 | False | 0.012028 | 1.123430 | 1.000000 |
| 3 | GalOS(b1-3)[Neu5Ac(a2-6)]GalNAc | 2.118119 | 0.270345 | 1.545193e-01 | 1.000000e+00 | False | 0.006427 | 0.524108 | 0.857944 |
Okay, seems like there are quite a few glycan that are either higher or lower in the cancer samples (see the “Log2FC” or “Effect size” columns for the direction of the effect)! A really important column here is the “corr p-val” column, which indicates the p-values of the comparisons after a two-stage Benjamini-Hochberg correction for multiple testing. By comparing this with a significance threshold, we can tell whether the observed difference is statistically significant. Usually, this threshold is p < 0.05 but this is a bad convention, since the meaning of p-values changes with sample size (a p-value of 0.049 at a sample size of 1,000 should not be considered significant but it should at a sample size of 10). glycowork adjusts this for you.
If you want to know what’s happening under the hood, this function is transforming your data via a CLR-transformation with a scale uncertainty model to counteract (i) the compositional nature of glycomics data and (ii) potential absolute differences between your conditions. In any case, let’s now have a look at motifs/substructures! While you can further specify which motifs you want to look at via the “feature_set” option, simply switching on “motifs = True” can be sufficient to now analyze at the motif level.
df_res = get_differential_expression(df, [1,3,5,7,9,11,13,15,17], [2,4,6,8,10,12,14,16,18], paired = True, motifs = True)
df_resYou're working with an alpha of 0.05632533070968504 that has been adjusted for your sample size of 18.| Glycan | Mean abundance | Log2FC | p-val | corr p-val | significant | corr Levene p-val | Effect size | Equivalence p-val | |
|---|---|---|---|---|---|---|---|---|---|
| 14 | Gal(b1-3)GalNAc | 11.619452 | -2.063177 | 2.439948e-13 | 4.879895e-13 | True | 0.013898 | -30.228627 | 1.00000 |
| 10 | Neu5Ac | 18.852289 | -2.787844 | 1.040438e-11 | 1.040438e-11 | True | 0.001238 | -18.897011 | 1.00000 |
| 6 | Gal | 16.868429 | -2.516696 | 1.133056e-11 | 1.133056e-11 | True | 0.020255 | -18.696195 | 1.00000 |
| 12 | Neu5Ac(a2-3)Gal | 13.543725 | -2.236032 | 2.550679e-11 | 2.550679e-11 | True | 0.002325 | -16.888493 | 1.00000 |
| 7 | GalNAc | 11.873553 | -2.081756 | 3.852593e-11 | 3.852593e-11 | True | 0.008764 | -16.037534 | 1.00000 |
| 0 | H_antigen_type2 | 0.205808 | 4.268800 | 3.465405e-09 | 3.465405e-09 | True | 0.002325 | 9.109177 | 1.00000 |
| 13 | Neu5Ac(a2-8)Neu5Ac | 0.102184 | 4.495409 | 4.208184e-09 | 4.208184e-09 | True | 0.006341 | 8.888582 | 1.00000 |
| 4 | Oglycan_core1 | 7.947107 | -1.625970 | 8.225545e-09 | 8.225545e-09 | True | 0.013050 | -8.166720 | 1.00000 |
| 3 | Disialyl_T_antigen | 4.859749 | -0.974023 | 1.321779e-07 | 1.321779e-07 | True | 0.016687 | -5.736893 | 1.00000 |
| 8 | GalOS | 0.254101 | 2.927287 | 2.127082e-07 | 2.127082e-07 | True | 0.002342 | 5.397373 | 1.00000 |
| 11 | Neu5Ac(a2-6)GalNAc | 4.952280 | -0.954426 | 2.642801e-07 | 2.642801e-07 | True | 0.010189 | -5.248866 | 1.00000 |
| 2 | Terminal_LacNAc_type2 | 0.483677 | 2.482075 | 5.753778e-07 | 5.753778e-07 | True | 0.009322 | 4.747614 | 1.00000 |
| 9 | GlcNAc6S(b1-6)GalNAc | 1.576631 | 1.105146 | 4.584820e-05 | 4.584820e-05 | True | 0.004694 | 2.648669 | 1.00000 |
| 5 | Mucin_elongated_core2 | 3.672346 | -0.149060 | 2.484354e-01 | 2.484354e-01 | False | 0.013898 | -0.414931 | 0.78216 |
| 1 | Internal_LacNAc_type2 | 3.188669 | 0.078740 | 3.643926e-01 | 3.643926e-01 | False | 0.012305 | 0.320546 | 0.78216 |
So here we see now that extended core 2 structures, especially sulfated ones, are significantly increasing in cancer, whereas core 1 structures do not change significantly. What’s interesting here is that most forms of sialylation are in fact decreasing in cancer, despite the typical finding that sialylation increases in tumor and despite the fact that there are individual sialylated structures that significantly increase. On the one hand, this of course depends on the tumor / type of cancer. But it’s very important to keep that this is precisely why it’s key to analyze glycomics data at the motif level to get an overall picture, as the intuition can often be led astray by looking at individual structures.
Okay, last but not least, let’s make a volcano plot with the results we got from the motif-level analysis! For this, we can use the glycowork.motif.analysis.get_volcano function, which takes the output from glycowork.motif.analysis.get_differential_expression and can also be further styled with GlycoDraw as is discussed below.
get_volcano(df_res, n = 18)You're working with an alpha of 0.05632533070968504 that has been adjusted for your sample size of 18.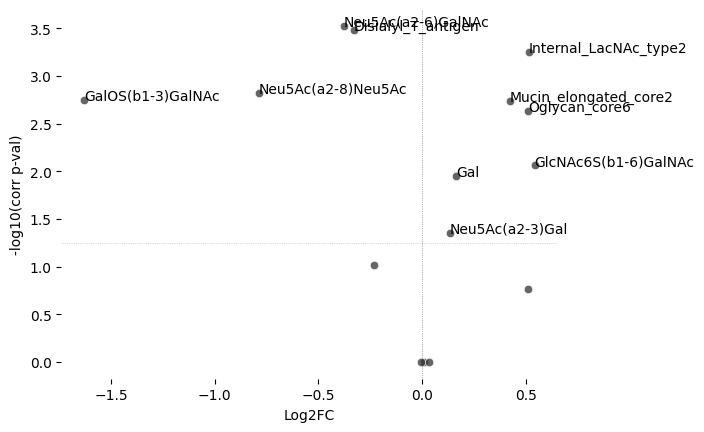
Deep Learning Code Snippets
For those of you that are blessed with GPU-access, we wrote up a minimal example for training a glycan-focused deep learning model. Even if you usually do not have access to a GPU, you can always go to Google Colab (click on the badge at the top of this site to open this notebook on Google Colab) and change the runtime type to GPU. Then you can paste the following code in a cell, execute, and see our model train!
First, we need to install torch_geometric for deep learning (ideally keep the package versions; especially if you want to use our trained models):
!pip install torch==2.5.1
!pip install torch-geometric==2.6.0Alternatively, if you’ve installed glycowork like this, you likely have everything in place already:
pip install glycowork[ml]SweetNet-type model
Then, we import the relevant glycowork modules:
from glycowork.glycan_data.loader import df_species
from glycowork.ml.train_test_split import hierarchy_filter
from glycowork.ml.processing import split_data_to_train
from glycowork.ml import models
from glycowork.ml import model_trainingFinally, we can prepare the data and train a model to classify from which taxonomic kingdom a glycan came from, using our graph convolutional neural network, SweetNet.
train_x, val_x, train_y, val_y, id_val, class_list, class_converter = hierarchy_filter(df_species,
rank = 'Kingdom')
dataloaders = split_data_to_train(train_x, val_x, train_y, val_y)
model = models.prep_model('SweetNet', len(class_list))
optimizer_ft, scheduler, criterion = model_training.training_setup(model, 0.0005, num_classes = len(class_list))
model_ft = model_training.train_model(model, dataloaders, criterion, optimizer_ft, scheduler,
num_epochs = 100)And that’s it! You’ve just trained a state-of-the-art deep learning model with effectively five lines of code (excluding package imports of course).
LectinOracle-type model
For LectinOracle, you need a protein sequence and glycan sequences. Further, you will need to retrieve the ESMC-representation for your protein sequence, as that is used as the actual input for LectinOracle. If you don’t want to do that, check below for the LectinOracle_flex model. Here the minimal process is shown for the arbitrary protein sequence “QWERTFVCF”. The trained model is yet again retrieved via prep_model.
!pip install fair-esm
from esm.models.esmc import ESMC
from glycowork.ml.inference import get_esmc_representations, get_lectin_preds
from glycowork.ml.models import prep_model
model = ESMC.from_pretrained("esmc_300m")
rep = get_esmc_representations(["QWERTFVCF"], model)
leor = prep_model("LectinOracle", 1, trained = True)
get_lectin_preds("QWERTFVCF", ['Neu5Ac(a2-6)GalNAc', 'Gal(b1-4)Glc'], leor, rep)The output here (same as for LectinOracle_flex below) is a table/dataframe, where each row shows the tested glycan and its associated binding prediction. The higher the binding prediction, the more likely an interaction (negative values in general likely mean “no interaction”), with a moderate correlation with binding affinity (based on our findings so far).
LectinOracle_flex-type model
While it has a slightly lower performance than LectinOracle, LectinOracle_flex can directly use protein sequences as input (no ESMC-conversion necessary). This is made possible by knowledge distillation, in which we train a sub-module to mimic the conversion of the much larger ESMC model. Just get the correct model via prep_model and declare flex = True and you’re good to go!
from glycowork.ml.inference import get_lectin_preds
from glycowork.ml.models import prep_model
leor = prep_model("LectinOracle_flex", 1, trained = True)
get_lectin_preds("QWERTFVCF", ['Neu5Ac(a2-6)GalNAc', 'Gal(b1-4)Glc'], leor, flex = True)GlycoDraw Code Snippets
GlycoDraw('Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc')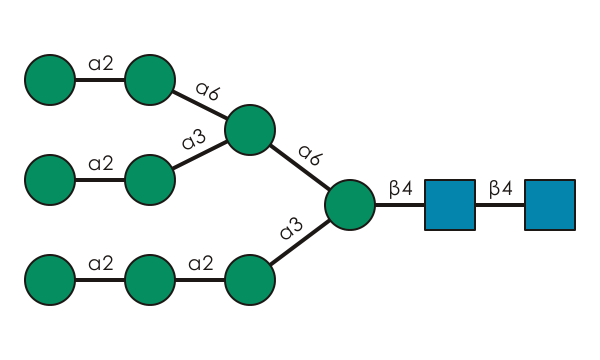
Okay, that’s all nice and well. But what if you don’t have lots of space in your figure?
Enter compact = True
GlycoDraw('Man(a1-2)Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-3)[Man(a1-2)Man(a1-6)]Man(a1-6)]Man(b1-4)GlcNAc(b1-4)GlcNAc',
compact = True)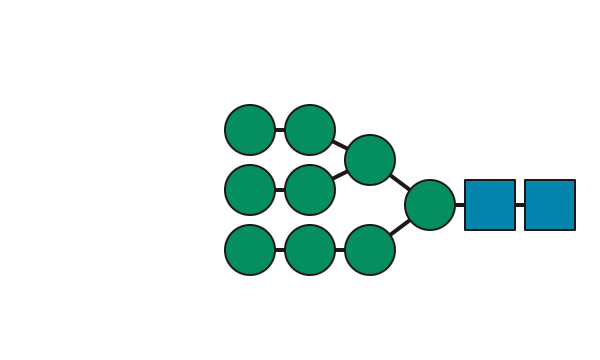
You can also go beyond mere drawing and add some more information, such as motif highlighting! In the below example, we highlight all Lewis antigen motifs in that structure.
GlycoDraw('Neu5Ac(a2-3)Gal(b1-4)[Fuc(a1-3)]GlcNAc(b1-2)Man(a1-3)[Gal(b1-3)[Fuc(a1-4)]GlcNAc(b1-2)Man(a1-6)]Man(b1-4)GlcNAc(b1-4)[Fuc(a1-6)]GlcNAc',
highlight_motif = 'Fuc(a1-?)[Gal(b1-?)]GlcNAc')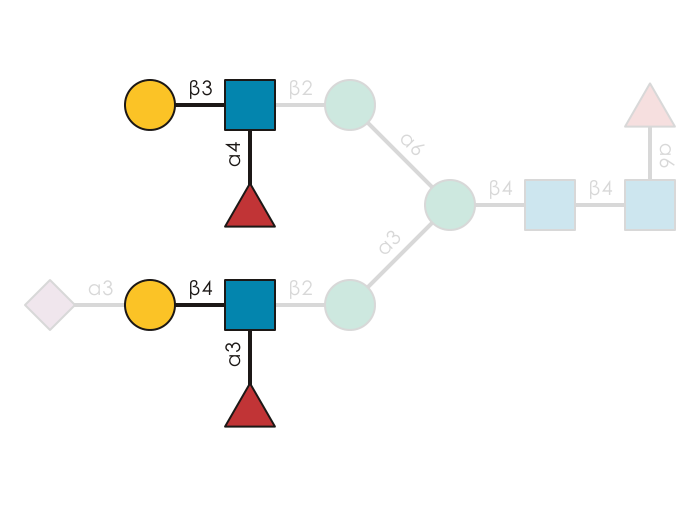
Or you could draw repeat structures, such as in glycosaminoglycans (GAGs). Here’s one example:
GlycoDraw('GalNAc4S(b1-4)GlcA(b1-3)', repeat = '2-3')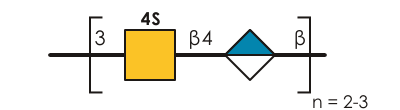
Or how about visualizing flexibility, attention, or any kind of attribution on a per-monosaccharide basis?
GlycoDraw("Fuc(a1-2)[Neu5Ac(a2-3)]Gal(b1-3)[Neu5Ac(a2-8)Neu5Ac(a2-6)]GalNAc", per_residue = [0.9, 0.7, 0.3, 0.2, 0.5, 0.1])
And that’s only a brief glimpse at the offered functionality! Why not replace the text labels from glycowork.motif.analysis.get_heatmap or glycowork.motif.analysis.get_volcano with glycan images via glycowork.motif.draw.annotate_figure?
Be sure to check out Lundstrøm et al., Glycobiology, 2023 for more details and inspirations!